Spatial and temporal organization of chromosome duplication and segregation in the cyanobacterium Synechococcus elongatus PCC 7942
- PMID: 23112856
- PMCID: PMC3480399
- DOI: 10.1371/journal.pone.0047837
Spatial and temporal organization of chromosome duplication and segregation in the cyanobacterium Synechococcus elongatus PCC 7942
Abstract
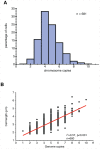
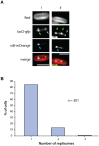
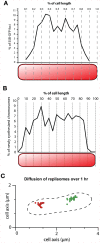
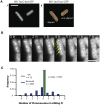
The spatial and temporal control of chromosome duplication and segregation is crucial for proper cell division. While this process is well studied in eukaryotic and some prokaryotic organisms, relatively little is known about it in prokaryotic polyploids such as Synechococcus elongatus PCC 7942, which is known to possess one to eight copies of its single chromosome. Using a fluorescent repressor-operator system, S. elongatus chromosomes and chromosome replication forks were tagged and visualized. We found that chromosomal duplication is asynchronous and that the total number of chromosomes is correlated with cell length. Thus, replication is independent of cell cycle and coupled to cell growth. Replication events occur in a spatially random fashion. However, once assembled, replisomes move in a constrained manner. On the other hand, we found that segregation displays a striking spatial organization in some cells. Chromosomes transiently align along the major axis of the cell and timing of alignment was correlated to cell division. This mechanism likely contributes to the non-random segregation of chromosome copies to daughter cells.
Conflict of interest statement
Figures

Similar articles
-
Spatial ordering of chromosomes enhances the fidelity of chromosome partitioning in cyanobacteria.Proc Natl Acad Sci U S A. 2012 Aug 21;109(34):13638-43. doi: 10.1073/pnas.1211144109. Epub 2012 Aug 6. Proc Natl Acad Sci U S A. 2012. PMID: 22869746 Free PMC article.
-
ParA-like protein influences the distribution of multi-copy chromosomes in cyanobacterium Synechococcus elongatus PCC 7942.Microbiology (Reading). 2018 Jan;164(1):45-56. doi: 10.1099/mic.0.000577. Epub 2017 Nov 21. Microbiology (Reading). 2018. PMID: 29165230
-
Intensive DNA Replication and Metabolism during the Lag Phase in Cyanobacteria.PLoS One. 2015 Sep 2;10(9):e0136800. doi: 10.1371/journal.pone.0136800. eCollection 2015. PLoS One. 2015. PMID: 26331851 Free PMC article.
-
Cyanobacterial multi-copy chromosomes and their replication.Biosci Biotechnol Biochem. 2020 Jul;84(7):1309-1321. doi: 10.1080/09168451.2020.1736983. Epub 2020 Mar 11. Biosci Biotechnol Biochem. 2020. PMID: 32157949 Review.
-
The chromosome cycle of prokaryotes.Mol Microbiol. 2013 Oct;90(2):214-27. doi: 10.1111/mmi.12372. Epub 2013 Sep 8. Mol Microbiol. 2013. PMID: 23962352 Free PMC article. Review.
Cited by
-
The McdAB system positions α-carboxysomes in proteobacteria.Mol Microbiol. 2021 Jul;116(1):277-297. doi: 10.1111/mmi.14708. Epub 2021 Mar 8. Mol Microbiol. 2021. PMID: 33638215 Free PMC article.
-
Discrete gene replication events drive coupling between the cell cycle and circadian clocks.Proc Natl Acad Sci U S A. 2016 Apr 12;113(15):4063-8. doi: 10.1073/pnas.1507291113. Epub 2016 Mar 28. Proc Natl Acad Sci U S A. 2016. PMID: 27035936 Free PMC article.
-
Live-cell imaging of cyanobacteria.Photosynth Res. 2015 Oct;126(1):33-46. doi: 10.1007/s11120-014-0049-x. Epub 2014 Nov 4. Photosynth Res. 2015. PMID: 25366827 Review.
-
The allometry of cellular DNA and ribosomal gene content among microbes and its use for the assessment of microbiome community structure.Microbiome. 2021 Aug 17;9(1):173. doi: 10.1186/s40168-021-01111-z. Microbiome. 2021. PMID: 34404486 Free PMC article.
-
Bacterial Argonaute Proteins Aid Cell Division in the Presence of Topoisomerase Inhibitors in Escherichia coli.Microbiol Spectr. 2023 Jun 15;11(3):e0414622. doi: 10.1128/spectrum.04146-22. Epub 2023 Apr 27. Microbiol Spectr. 2023. PMID: 37102866 Free PMC article.
References
-
- Lau IF, Filipe SR, Søballe B, Økstad O-A, Barre F-X, et al.. (2003) Spatial and temporal organization of replicating Escherichia coli chromosomes. Molecular microbiology 49: 731–743. Available: http://www.ncbi.nlm.nih.gov/pubmed/12864855. Accessed 2012 Jul 13. - PubMed
-
- Berlatzky IA, Rouvinski A, Ben-Yehuda S (2008) Spatial organization of a replicating bacterial chromosome. Proceedings of the National Academy of Sciences of the United States of America 105: 14136–14140. Available: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2544591&tool=p.... Accessed 2012 Jul 13. - PMC - PubMed
-
- Toro E, Shapiro L (2010) Bacterial chromosome organization and segregation. Cold Spring Harbor perspectives in biology 2: a000349. Available: http://cshperspectives.cshlp.org/content/2/2/a000349.full. Accessed 2012 Jul 16. - PMC - PubMed
-
- Graumann PL, Dame RT, Dorman CJ (2010) The Chromosome Segregation Machinery in Bacteria. Bacterial Chromatin (Google eBook). Springer. 31–48. Available: http://books.google.com/books?id=Esi4D4NjvskC&pgis=1. Accessed 2012 Jul 13.
-
- Haeusser DP, Levin PA (2008) The great divide: coordinating cell cycle events during bacterial growth and division. Current opinion in microbiology 11: 94–99. Available: http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2397022&tool=p.... Accessed 2012 Jul 13. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

