Multiple NF-κB sites in HIV-1 subtype C long terminal repeat confer superior magnitude of transcription and thereby the enhanced viral predominance
- PMID: 23132857
- PMCID: PMC3531786
- DOI: 10.1074/jbc.M112.397158
Multiple NF-κB sites in HIV-1 subtype C long terminal repeat confer superior magnitude of transcription and thereby the enhanced viral predominance
Abstract
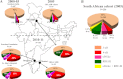
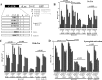
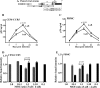
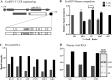
We demonstrate that at least three different promoter variant strains of HIV-1 subtype C have been gradually expanding and replacing the standard subtype C viruses in India, and possibly in South Africa and other global regions, over the past decade. The new viral strains contain an additional NF-κB, NF-κB-like, or RBEIII site in the viral promoter. Although the acquisition of an additional RBEIII site is a property shared by all the HIV-1 subtypes, acquiring an additional NF-κB site remains an exclusive property of subtype C. The acquired κB site is genetically distinct, binds the p50-p65 heterodimer, and strengthens the viral promoter at the levels of transcription initiation and elongation. The 4-κB viruses dominate the 3-κB "isogenic" viral strains in pairwise competition assays in T-cell lines, primary cells, and the ecotropic human immunodeficiency virus mouse model. The dominance of the 4-κB viral strains is also evident in the natural context when the subjects are coinfected with κB-variant viral strains. The mean plasma viral loads, but not CD4 counts, are significantly different in 4-κB infection suggesting that these newly emerging strains are probably more infectious. It is possible that higher plasma viral loads underlie selective transmission of the 4-κB viral strains. Several publications previously reported duplication or deletion of diverse transcription factor-binding sites in the viral promoter. Unlike previous reports, our study provides experimental evidence that the new viral strains gained a potential selective advantage as a consequence of the acquired transcription factor-binding sites and importantly that these strains have been expanding at the population level.
Figures


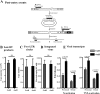

References
-
- Tebit D. M., Arts E. J. (2011) Tracking a century of global expansion and evolution of HIV to drive understanding and to combat disease. Lancet Infect. Dis. 11, 45–56 - PubMed
-
- Bongertz V., Bou-Habib D. C., Brígido L. F., Caseiro M., Chequer P. J., Couto-Fernandez J. C., Ferreira P. C., Galvão-Castro B., Greco D., Guimarães M. L., Linhares de Carvalho M. I., Morgado M. G., Oliveira C. A., Osmanov S., Ramos C. A., Rossini M., Sabino E., Tanuri A., Ueda M. (2000) HIV-1 diversity in Brazil. Genetic, biologic, and immunologic characterization of HIV-1 strains in three potential HIV vaccine evaluation sites. Brazilian Network for HIV Isolation and Characterization. J. Acquir. Immune. Defic. Syndr. 23, 184–193 - PubMed
-
- Soares M. A., De Oliveira T., Brindeiro R. M., Diaz R. S., Sabino E. C., Brigido L., Pires I. L., Morgado M. G., Dantas M. C., Barreira D., Teixeira P. R., Cassol S., Tanuri A. (2003) A specific subtype C of human immunodeficiency virus type 1 circulates in Brazil. AIDS 17, 11–21 - PubMed
-
- Locateli D., Stoco P. H., de Queiroz A. T., Alcântara L. C., Ferreira L. G., Zanetti C. R., Rodrigues R., Grisard E. C., Pinto A. R. (2007) Molecular epidemiology of HIV-1 in Santa Catarina State confirms increases of subtype C in Southern Brazil. J. Med. Virol. 79, 1455–1463 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

