Metagenomic analysis of the microbiota from the crop of an invasive snail reveals a rich reservoir of novel genes
- PMID: 23133637
- PMCID: PMC3486852
- DOI: 10.1371/journal.pone.0048505
Metagenomic analysis of the microbiota from the crop of an invasive snail reveals a rich reservoir of novel genes
Abstract
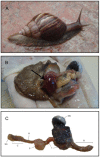
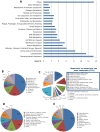
The shortage of petroleum reserves and the increase in CO(2) emissions have raised global concerns and highlighted the importance of adopting sustainable energy sources. Second-generation ethanol made from lignocellulosic materials is considered to be one of the most promising fuels for vehicles. The giant snail Achatina fulica is an agricultural pest whose biotechnological potential has been largely untested. Here, the composition of the microbial population within the crop of this invasive land snail, as well as key genes involved in various biochemical pathways, have been explored for the first time. In a high-throughput approach, 318 Mbp of 454-Titanium shotgun metagenomic sequencing data were obtained. The predominant bacterial phylum found was Proteobacteria, followed by Bacteroidetes and Firmicutes. Viruses, Fungi, and Archaea were present to lesser extents. The functional analysis reveals a variety of microbial genes that could assist the host in the degradation of recalcitrant lignocellulose, detoxification of xenobiotics, and synthesis of essential amino acids and vitamins, contributing to the adaptability and wide-ranging diet of this snail. More than 2,700 genes encoding glycoside hydrolase (GH) domains and carbohydrate-binding modules were detected. When we compared GH profiles, we found an abundance of sequences coding for oligosaccharide-degrading enzymes (36%), very similar to those from wallabies and giant pandas, as well as many novel cellulase and hemicellulase coding sequences, which points to this model as a remarkable potential source of enzymes for the biofuel industry. Furthermore, this work is a major step toward the understanding of the unique genetic profile of the land snail holobiont.
Conflict of interest statement
Figures


Similar articles
-
Metagenomic analysis of the Rhinopithecus bieti fecal microbiome reveals a broad diversity of bacterial and glycoside hydrolase profiles related to lignocellulose degradation.BMC Genomics. 2015 Mar 12;16(1):174. doi: 10.1186/s12864-015-1378-7. BMC Genomics. 2015. PMID: 25887697 Free PMC article.
-
Metagenomic insights into the fibrolytic microbiome in yak rumen.PLoS One. 2012;7(7):e40430. doi: 10.1371/journal.pone.0040430. Epub 2012 Jul 13. PLoS One. 2012. PMID: 22808161 Free PMC article.
-
Gut bacterial communities in the giant land snail Achatina fulica and their modification by sugarcane-based diet.PLoS One. 2012;7(3):e33440. doi: 10.1371/journal.pone.0033440. Epub 2012 Mar 15. PLoS One. 2012. PMID: 22438932 Free PMC article.
-
Engineering Ligninolytic Consortium for Bioconversion of Lignocelluloses to Ethanol and Chemicals.Protein Pept Lett. 2018;25(2):108-119. doi: 10.2174/0929866525666180122105835. Protein Pept Lett. 2018. PMID: 29359652 Review.
-
A review of biological delignification and detoxification methods for lignocellulosic bioethanol production.Crit Rev Biotechnol. 2015;35(3):342-54. doi: 10.3109/07388551.2013.878896. Crit Rev Biotechnol. 2015. PMID: 24506661 Review.
Cited by
-
Environment and Diet Influence the Bacterial Microbiome of Ambigolimax valentianus, an Invasive Slug in California.Insects. 2021 Jun 23;12(7):575. doi: 10.3390/insects12070575. Insects. 2021. PMID: 34201881 Free PMC article.
-
Role of metagenomics in prospecting novel endoglucanases, accentuating functional metagenomics approach in second-generation biofuel production: a review.Biomass Convers Biorefin. 2023;13(2):1371-1398. doi: 10.1007/s13399-020-01186-y. Epub 2021 Jan 7. Biomass Convers Biorefin. 2023. PMID: 33437563 Free PMC article. Review.
-
The Microbiome of Animals: Implications for Conservation Biology.Int J Genomics. 2016;2016:5304028. doi: 10.1155/2016/5304028. Epub 2016 Apr 18. Int J Genomics. 2016. PMID: 27195280 Free PMC article. Review.
-
Metagenomic Analysis of the Gut Microbiome of the Common Black Slug Arion ater in Search of Novel Lignocellulose Degrading Enzymes.Front Microbiol. 2017 Nov 8;8:2181. doi: 10.3389/fmicb.2017.02181. eCollection 2017. Front Microbiol. 2017. PMID: 29167663 Free PMC article.
-
Mollusk microbiota shift during Angiostrongylus cantonensis infection in the freshwater snail Biomphalaria glabrata and the terrestrial slug Phillocaulis soleiformis.Parasitol Res. 2020 Aug;119(8):2495-2503. doi: 10.1007/s00436-020-06743-y. Epub 2020 Jun 17. Parasitol Res. 2020. PMID: 32556501
References
-
- Goldemberg J (2007) Ethanol for a sustainable energy future. Science 315: 808–810. - PubMed
-
- Somerville C, Youngs H, Taylor C, Davis SC, Long SP (2010) Feedstocks for Lignocellulosic Biofuels. Science 329: 790–792. - PubMed
-
- Jenkins BM, Baxter LL, Miles-Jr TR, Miles TR (1998) Combustion properties of biomass. Fuel Proc Technol 54: 17–46.
-
- Ragauskas AJ, Williams CK, Davison BH, Britovsek G, Cairney J, et al. (2006) The Path Forward for Biofuels and Biomaterials. Science 311: 484–489. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

