Maternally recruited DCP1A and DCP2 contribute to messenger RNA degradation during oocyte maturation and genome activation in mouse
- PMID: 23136299
- PMCID: PMC4434936
- DOI: 10.1095/biolreprod.112.105312
Maternally recruited DCP1A and DCP2 contribute to messenger RNA degradation during oocyte maturation and genome activation in mouse
Abstract
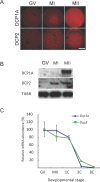
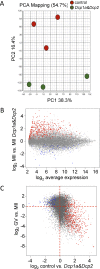
The oocyte-to-zygote transition entails transforming a highly differentiated oocyte into totipotent blastomeres and represents one of the earliest obstacles that must be successfully hurdled for continued development. Degradation of maternal mRNAs, which likely lies at the heart of this transition, is characterized by a transition from mRNA stability to instability during oocyte maturation. Although phosphorylation of the oocyte-specific RNA-binding protein MSY2 during maturation is implicated in making maternal mRNAs more susceptible to degradation, mechanisms underlying mRNA degradation during oocyte maturation remain poorly understood. We report that DCP1A and DCP2, proteins responsible for decapping mRNA, are encoded by maternal mRNAs recruited for translation during maturation via cytoplasmic polyadenylation elements located in their 3' untranslated regions. Both DCP1A and DCP2 are phosphorylated during maturation, with CDC2A being the kinase likely responsible for both, although MAPK may be involved in DCP1A phosphorylation. Inhibiting accumulation of DCP1A and DCP2 by RNA interference or morpholinos decreases not only degradation of mRNAs during meiotic maturation but also transcription of the zygotic genome. The results indicate that maternally recruited DCP1A and DCP2 are critical players in the transition from mRNA stability to instability during meiotic maturation and that proper maternal mRNA degradation must be successful to execute the oocyte-to-zygote transition.
Figures





References
-
- Tucker M, Valencia-Sanchez MA, Staples RR, Chen J, Denis CL, Parker R. The transcription factor associated Ccr4 and Caf1 proteins are components of the major cytoplasmic mRNA deadenylase in Saccharomyces cerevisiae. Cell. 2001;104:377–386. - PubMed
-
- Coller J, Parker R. Eukaryotic mRNA decapping. Annu Rev Biochem. 2004;73:861–890. - PubMed
-
- Beelman CA, Stevens A, Caponigro G, LaGrandeur TE, Hatfield L, Fortner DM, Parker R. An essential component of the decapping enzyme required for normal rates of mRNA turnover. Nature. 1996;382:642–646. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

