Genome-wide control of polyadenylation site choice by CPSF30 in Arabidopsis
- PMID: 23136375
- PMCID: PMC3531840
- DOI: 10.1105/tpc.112.096107
Genome-wide control of polyadenylation site choice by CPSF30 in Arabidopsis
Abstract
The Arabidopsis thaliana ortholog of the 30-kD subunit of the mammalian Cleavage and Polyadenylation Specificity Factor (CPSF30) has been implicated in the responses of plants to oxidative stress, suggesting a role for alternative polyadenylation. To better understand this, poly(A) site choice was studied in a mutant (oxt6) deficient in CPSF30 expression using a genome-scale approach. The results indicate that poly(A) site choice in a large majority of Arabidopsis genes is altered in the oxt6 mutant. A number of poly(A) sites were identified that are seen only in the wild type or oxt6 mutant. Interestingly, putative polyadenylation signals associated with sites that are seen only in the oxt6 mutant are decidedly different from the canonical plant polyadenylation signal, lacking the characteristic A-rich near-upstream element (where AAUAAA can be found); this suggests that CPSF30 functions in the handling of the near-upstream element. The sets of genes that possess sites seen only in the wild type or mutant were enriched for those involved in stress and defense responses, a result consistent with the properties of the oxt6 mutant. Taken together, these studies provide new insights into the mechanisms and consequences of CPSF30-mediated alternative polyadenylation.
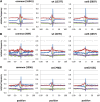
Figures
Similar articles
-
Revisiting CPSF30-mediated alternative polyadenylation in Arabidopsis thaliana.PLoS One. 2025 Feb 24;20(2):e0319180. doi: 10.1371/journal.pone.0319180. eCollection 2025. PLoS One. 2025. PMID: 39992955 Free PMC article.
-
Integration of developmental and environmental signals via a polyadenylation factor in Arabidopsis.PLoS One. 2014 Dec 29;9(12):e115779. doi: 10.1371/journal.pone.0115779. eCollection 2014. PLoS One. 2014. PMID: 25546057 Free PMC article.
-
Transcriptome Analyses of FY Mutants Reveal Its Role in mRNA Alternative Polyadenylation.Plant Cell. 2019 Oct;31(10):2332-2352. doi: 10.1105/tpc.18.00545. Epub 2019 Aug 19. Plant Cell. 2019. PMID: 31427469 Free PMC article.
-
CPSF30 at the Interface of Alternative Polyadenylation and Cellular Signaling in Plants.Biomolecules. 2015 Jun 8;5(2):1151-68. doi: 10.3390/biom5021151. Biomolecules. 2015. PMID: 26061761 Free PMC article. Review.
-
The Arabidopsis polyadenylation factor subunit CPSF30 as conceptual link between mRNA polyadenylation and cellular signaling.Curr Opin Plant Biol. 2014 Oct;21:128-132. doi: 10.1016/j.pbi.2014.07.002. Epub 2014 Aug 9. Curr Opin Plant Biol. 2014. PMID: 25104048 Review.
Cited by
-
Dynamics of alternative polyadenylation in single root cells of Arabidopsis thaliana.Front Plant Sci. 2024 Sep 20;15:1437118. doi: 10.3389/fpls.2024.1437118. eCollection 2024. Front Plant Sci. 2024. PMID: 39372861 Free PMC article.
-
Alternative polyadenylation profiles of susceptible and resistant rice (Oryza sativa L.) in response to bacterial leaf blight using RNA-seq.BMC Plant Biol. 2024 Feb 28;24(1):145. doi: 10.1186/s12870-024-04839-6. BMC Plant Biol. 2024. PMID: 38413866 Free PMC article.
-
Interplay between Alternative Splicing and Alternative Polyadenylation Defines the Expression Outcome of the Plant Unique OXIDATIVE TOLERANT-6 Gene.Sci Rep. 2017 May 17;7(1):2052. doi: 10.1038/s41598-017-02215-z. Sci Rep. 2017. PMID: 28515442 Free PMC article.
-
Transcript-level expression control of plant NLR genes.Mol Plant Pathol. 2018 May;19(5):1267-1281. doi: 10.1111/mpp.12607. Epub 2017 Nov 20. Mol Plant Pathol. 2018. PMID: 28834153 Free PMC article. Review.
-
High throughput characterizations of poly(A) site choice in plants.Methods. 2014 May 1;67(1):74-83. doi: 10.1016/j.ymeth.2013.06.037. Epub 2013 Jul 11. Methods. 2014. PMID: 23851255 Free PMC article.
References
-
- Addepalli B., Hunt A.G. (2008). Redox and heavy metal effects on the biochemical activities of an Arabidopsis polyadenylation factor subunit. Arch. Biochem. Biophys. 473: 88–95 - PubMed
-
- Addepalli B., Limbach P.A., Hunt A.G. (2010). A disulfide linkage in a CCCH zinc finger motif of an Arabidopsis CPSF30 ortholog. FEBS Lett. 584: 4408–4412 - PubMed
-
- Barabino S.M., Hübner W., Jenny A., Minvielle-Sebastia L., Keller W. (1997). The 30-kD subunit of mammalian cleavage and polyadenylation specificity factor and its yeast homolog are RNA-binding zinc finger proteins. Genes Dev. 11: 1703–1716 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

