Expression and promoter analysis of a highly restricted integrin alpha gene in vascular smooth muscle
- PMID: 23142384
- PMCID: PMC3517168
- DOI: 10.1016/j.gene.2012.10.073
Expression and promoter analysis of a highly restricted integrin alpha gene in vascular smooth muscle
Abstract
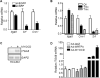
Full genome annotation requires gene expression analysis and elucidation of promoter activity. Here, we analyzed the expression and promoter of a highly restricted integrin gene, Itga8. RNase protection and quantitative RT-PCR showed Itga8 to be expressed most abundantly in vascular smooth muscle cells (SMC). Transcription start site mapping of Itga8 revealed the immediate 5' promoter region to be poorly conserved with orthologous sequences in the human genome. Further comparative sequence analysis showed a number of conserved non-coding sequence modules around the Itga8 gene. The immediate promoter region and an upstream conserved sequence module were each found to contain a CArG box, which is a binding site for serum response factor (SRF). Luciferase reporter assays revealed activity of several Itga8 promoter constructs with no apparent restricted activity to SMC types. Further, neither SRF nor its coactivator, Myocardin (MYOCD), was able to induce several distinct Itga8 promoter constructs. Transgenic mouse studies failed to reveal Itga8 promoter activity, indicating distal regulatory elements likely control this gene's in vivo expression profile. Interestingly, although the promoter was unresponsive to SRF/MYOCD, the endogenous Itga8 gene showed increases in expression upon ectopic MYOCD expression even though knockdown of SRF both in vitro and in vivo failed to demonstrate a corresponding change in Itga8. Collectively, these data demonstrate that Itga8 expression is CArG-SRF independent, but MYOCD dependent through an as yet unknown sequence module that is distal from the promoter region.
Copyright © 2012 Elsevier B.V. All rights reserved.
Figures





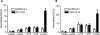
Similar articles
-
The smooth muscle cell-restricted KCNMB1 ion channel subunit is a direct transcriptional target of serum response factor and myocardin.J Biol Chem. 2009 Nov 27;284(48):33671-82. doi: 10.1074/jbc.M109.050419. Epub 2009 Oct 1. J Biol Chem. 2009. PMID: 19801679 Free PMC article.
-
Contribution of serum response factor and myocardin to transcriptional regulation of smoothelins.Cardiovasc Res. 2006 Apr 1;70(1):136-45. doi: 10.1016/j.cardiores.2005.12.018. Epub 2006 Jan 31. Cardiovasc Res. 2006. PMID: 16451796
-
Coronary Disease-Associated Gene TCF21 Inhibits Smooth Muscle Cell Differentiation by Blocking the Myocardin-Serum Response Factor Pathway.Circ Res. 2020 Feb 14;126(4):517-529. doi: 10.1161/CIRCRESAHA.119.315968. Epub 2019 Dec 9. Circ Res. 2020. PMID: 31815603 Free PMC article.
-
Serum response factor: toggling between disparate programs of gene expression.J Mol Cell Cardiol. 2003 Jun;35(6):577-93. doi: 10.1016/s0022-2828(03)00110-x. J Mol Cell Cardiol. 2003. PMID: 12788374 Review.
-
Myocardin: A novel player in atherosclerosis.Atherosclerosis. 2017 Feb;257:266-278. doi: 10.1016/j.atherosclerosis.2016.12.002. Epub 2016 Dec 1. Atherosclerosis. 2017. PMID: 28012646 Review.
Cited by
-
Inflammation and vascular smooth muscle cell dedifferentiation following carotid artery ligation.Physiol Genomics. 2017 Mar 1;49(3):115-126. doi: 10.1152/physiolgenomics.00095.2016. Epub 2016 Dec 30. Physiol Genomics. 2017. PMID: 28039430 Free PMC article.
-
Transcriptomic signatures of atheroresistance in the human atrium and ventricle highlight potential candidates for targeted atherosclerosis therapeutics.Biochem Biophys Rep. 2025 Apr 8;42:102007. doi: 10.1016/j.bbrep.2025.102007. eCollection 2025 Jun. Biochem Biophys Rep. 2025. PMID: 40248137 Free PMC article.
-
The short and long of noncoding sequences in the control of vascular cell phenotypes.Cell Mol Life Sci. 2015 Sep;72(18):3457-88. doi: 10.1007/s00018-015-1936-9. Epub 2015 May 29. Cell Mol Life Sci. 2015. PMID: 26022065 Free PMC article. Review.
-
Vascular smooth muscle-specific YAP/TAZ deletion triggers aneurysm development in mouse aorta.JCI Insight. 2023 Sep 8;8(17):e170845. doi: 10.1172/jci.insight.170845. JCI Insight. 2023. PMID: 37561588 Free PMC article.
-
Digital spatial profiling of segmental outflow regions in trabecular meshwork reveals a role for ADAM15.PLoS One. 2024 Feb 23;19(2):e0298802. doi: 10.1371/journal.pone.0298802. eCollection 2024. PLoS One. 2024. PMID: 38394161 Free PMC article.
References
-
- Chen J, Kitchen CM, Streb JW, Miano JM. Myocardin: a component of a molecular switch for smooth muscle differentiation. J. Mol. Cell. Cardiol. 2002;34:1345–1356. - PubMed
-
- Chen J, Maltby KM, Miano JM. A novel retinoid-response gene set in vascular smooth muscle cells. Biochem. Biophys. Res. Commun. 2001;281:475–482. - PubMed
-
- Ekwa-Ekoka C, Diaz GA, Carlson C, Hasegawa T, Samudrala R, Lim K-C, Yabu JM, Levy B, Schnapp LM. Genomic organization and sequence variation of the human integrin subunit α8 gene (ITGA8) Matrix Biol. 2004;23:487–496. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

