Predicting turns in proteins with a unified model
- PMID: 23144872
- PMCID: PMC3492357
- DOI: 10.1371/journal.pone.0048389
Predicting turns in proteins with a unified model
Abstract
Motivation: Turns are a critical element of the structure of a protein; turns play a crucial role in loops, folds, and interactions. Current prediction methods are well developed for the prediction of individual turn types, including α-turn, β-turn, and γ-turn, etc. However, for further protein structure and function prediction it is necessary to develop a uniform model that can accurately predict all types of turns simultaneously.
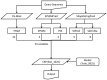
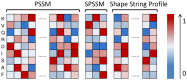
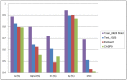
Results: In this study, we present a novel approach, TurnP, which offers the ability to investigate all the turns in a protein based on a unified model. The main characteristics of TurnP are: (i) using newly exploited features of structural evolution information (secondary structure and shape string of protein) based on structure homologies, (ii) considering all types of turns in a unified model, and (iii) practical capability of accurate prediction of all turns simultaneously for a query. TurnP utilizes predicted secondary structures and predicted shape strings, both of which have greater accuracy, based on innovative technologies which were both developed by our group. Then, sequence and structural evolution features, which are profile of sequence, profile of secondary structures and profile of shape strings are generated by sequence and structure alignment. When TurnP was validated on a non-redundant dataset (4,107 entries) by five-fold cross-validation, we achieved an accuracy of 88.8% and a sensitivity of 71.8%, which exceeded the most state-of-the-art predictors of certain type of turn. Newly determined sequences, the EVA and CASP9 datasets were used as independent tests and the results we achieved were outstanding for turn predictions and confirmed the good performance of TurnP for practical applications.
Conflict of interest statement
Figures

References
-
- Branden C, Tozz J (1991) Motifs of Protein Structure. Introduction to protein structure. 13–34.
-
- Meissner M, Koch O, Klebe G, Schneider G (2009) Prediction of turn types in protein structure by machine-learning classifiers. Proteins 74: 344–352. - PubMed
-
- Chou KC (2000) Prediction of tight turns and their types in proteins. Anal Biochem 286: 1–16. - PubMed
-
- Chou KC, Blinn JR (1997) Classification and prediction of beta-turn types. J Protein Chem 16: 575–595. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

