Identification of plasma lipid biomarkers for prostate cancer by lipidomics and bioinformatics
- PMID: 23152813
- PMCID: PMC3495963
- DOI: 10.1371/journal.pone.0048889
Identification of plasma lipid biomarkers for prostate cancer by lipidomics and bioinformatics
Abstract
Background: Lipids have critical functions in cellular energy storage, structure and signaling. Many individual lipid molecules have been associated with the evolution of prostate cancer; however, none of them has been approved to be used as a biomarker. The aim of this study is to identify lipid molecules from hundreds plasma apparent lipid species as biomarkers for diagnosis of prostate cancer.
Methodology/principal findings: Using lipidomics, lipid profiling of 390 individual apparent lipid species was performed on 141 plasma samples from 105 patients with prostate cancer and 36 male controls. High throughput data generated from lipidomics were analyzed using bioinformatic and statistical methods. From 390 apparent lipid species, 35 species were demonstrated to have potential in differentiation of prostate cancer. Within the 35 species, 12 were identified as individual plasma lipid biomarkers for diagnosis of prostate cancer with a sensitivity above 80%, specificity above 50% and accuracy above 80%. Using top 15 of 35 potential biomarkers together increased predictive power dramatically in diagnosis of prostate cancer with a sensitivity of 93.6%, specificity of 90.1% and accuracy of 97.3%. Principal component analysis (PCA) and hierarchical clustering analysis (HCA) demonstrated that patient and control populations were visually separated by identified lipid biomarkers. RandomForest and 10-fold cross validation analyses demonstrated that the identified lipid biomarkers were able to predict unknown populations accurately, and this was not influenced by patient's age and race. Three out of 13 lipid classes, phosphatidylethanolamine (PE), ether-linked phosphatidylethanolamine (ePE) and ether-linked phosphatidylcholine (ePC) could be considered as biomarkers in diagnosis of prostate cancer.
Conclusions/significance: Using lipidomics and bioinformatic and statistical methods, we have identified a few out of hundreds plasma apparent lipid molecular species as biomarkers for diagnosis of prostate cancer with a high sensitivity, specificity and accuracy.
Conflict of interest statement
Figures


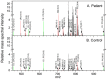
References
-
- Ramírez ML, Nelson EC, Evans CP (2008) Beyond prostate-specific antigen: alternate serum markers. Prostate Cancer Prostatic Dis 11 3 216–29. - PubMed
-
- Etzioni R, Penson DF, Legler JM, di Tommaso D, Boer R, et al. (2002) Over diagnosis due to prostate-specific antigen screening: lessons from U.S. prostate cancer incidence trends. Natl Cancer Inst 94 13 981–90. - PubMed
-
- Hamilton RJ, Platz EA, Freedland SJ (2009) Re: Prostate-specific antigen: a misused and maligned prostate cancer biomarker. J Natl Cancer Inst 101 8 611–2. - PubMed
-
- Elgamal AA, Holmes EH, Su SL, Tino WT, Simmons SJ, et al. (2000) Prostate-specific membrane antigen (PSMA): current benefits and future value. Semin Surg Oncol 18 1 10–6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

