Loss of the Par3 polarity protein promotes breast tumorigenesis and metastasis
- PMID: 23153534
- PMCID: PMC3500525
- DOI: 10.1016/j.ccr.2012.10.003
Loss of the Par3 polarity protein promotes breast tumorigenesis and metastasis
Erratum in
-
Loss of the Par3 Polarity Protein Promotes Breast Tumorigenesis and Metastasis.Cancer Cell. 2016 Aug 8;30(2):351-352. doi: 10.1016/j.ccell.2016.07.001. Epub 2016 Aug 8. Cancer Cell. 2016. PMID: 27505675 No abstract available.
Abstract
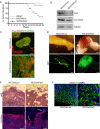
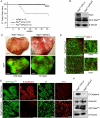
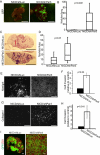
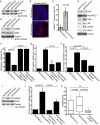
Loss of epithelial organization is a hallmark of carcinomas, but whether polarity regulates tumor growth and metastasis is poorly understood. To address this issue, we depleted the Par3 polarity gene by RNAi in combination with oncogenic Notch or Ras(61L) expression in the murine mammary gland. Par3 silencing dramatically reduced tumor latency in both models and produced invasive and metastatic tumors that retained epithelial marker expression. Par3 depletion was associated with induction of MMP9, destruction of the extracellular matrix, and invasion, all mediated by atypical PKC-dependant JAK/Stat3 activation. Importantly, Par3 expression is significantly reduced in human breast cancers, which correlates with active aPKC and Stat3. These data identify Par3 as a regulator of signaling pathways relevant to invasive breast cancer.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures



References
-
- Barbieri I, Pensa S, Pannellini T, Quaglino E, Maritano D, Demaria M, Voster A, Turkson J, Cavallo F, Watson CJ, et al. Constitutively active Stat3 enhances neu-mediated migration and metastasis in mammary tumors via upregulation of Cten. Cancer Res. 2010a;70:2558–2567. - PubMed
-
- Barbieri I, Quaglino E, Maritano D, Pannellini T, Riera L, Cavallo F, Forni G, Musiani P, Chiarle R, Poli V. Stat3 is required for anchorage-independent growth and metastasis but not for mammary tumor development downstream of the ErbB-2 oncogene. Mol Carcinog. 2010b;49:114–120. - PubMed
-
- Bilder D, Li M, Perrimon N. Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science. 2000;289:113–116. - PubMed
-
- Bromberg JF, Wrzeszczynska MH, Devgan G, Zhao Y, Pestell RG, Albanese C, Darnell JE., Jr. Stat3 as an oncogene. Cell. 1999;98:295–303. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

