Functional evolution of an anthocyanin pathway enzyme during a flower color transition
- PMID: 23155005
- PMCID: PMC3563968
- DOI: 10.1093/molbev/mss255
Functional evolution of an anthocyanin pathway enzyme during a flower color transition
Abstract
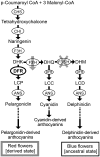
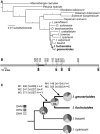
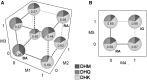
Dissecting the genetic basis for the evolution of species differences requires a combination of phylogenetic and molecular genetic perspectives. By mapping the genetic changes and their phenotypic effects onto the phylogeny, it is possible to distinguish changes that may have been directly responsible for a new character state from those that fine tune the transition. Here, we use phylogenetic and functional methods to trace the evolution of substrate specificity in dihydroflavonol-4-reductase (Dfr), an anthocyanin pathway gene known to be involved in the transition from blue to red flowers in Iochroma. Ancestral state reconstruction indicates that three substitutions occurred during the flower color transition, whereas several additional substitutions followed the transition. Comparisons of enzymatic function between ancestral proteins in blue- and red-flowered lineages and proteins from present-day taxa demonstrate that evolution of specificity for red pigment precursors was caused by the first three substitutions, which were fixed by positive selection and which differ from previously documented mutations affecting specificity. Two inferred substitutions subsequent to the initial flower color transition were also adaptive and resulted in an additional increase in specificity for red precursors. Epistatic interactions among both sets of substitutions may have limited the order of substitutions along branches of the phylogeny leading from blue-pigmented ancestors to the present-day red-flowered taxa. These results suggest that the species differences in DFR specificity may arise by a combination of selection on flower color and selection for improved pathway efficiency but that the exact series of genetic changes resulting in the evolution of specificity is likely to be highly contingent on the starting state.
Figures
Similar articles
-
Genetic basis for a rare floral mutant in an Andean species of Solanaceae.Am J Bot. 2015 Feb;102(2):264-72. doi: 10.3732/ajb.1400395. Epub 2015 Jan 22. Am J Bot. 2015. PMID: 25667079
-
Gene loss and parallel evolution contribute to species difference in flower color.Mol Biol Evol. 2011 Oct;28(10):2799-810. doi: 10.1093/molbev/msr109. Epub 2011 May 6. Mol Biol Evol. 2011. PMID: 21551271 Free PMC article.
-
Cloning and Functional Characterization of Dihydroflavonol 4-Reductase Gene Involved in Anthocyanidin Biosynthesis of Grape Hyacinth.Int J Mol Sci. 2019 Sep 24;20(19):4743. doi: 10.3390/ijms20194743. Int J Mol Sci. 2019. PMID: 31554290 Free PMC article.
-
Flower colour and cytochromes P450.Philos Trans R Soc Lond B Biol Sci. 2013 Jan 6;368(1612):20120432. doi: 10.1098/rstb.2012.0432. Print 2013 Feb 19. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 23297355 Free PMC article. Review.
-
Achievements and perspectives in biochemistry concerning anthocyanin modification for blue flower coloration.Plant Cell Physiol. 2015 Jan;56(1):28-40. doi: 10.1093/pcp/pcu097. Epub 2014 Jul 10. Plant Cell Physiol. 2015. PMID: 25015943 Review.
Cited by
-
Robustness of Reconstructed Ancestral Protein Functions to Statistical Uncertainty.Mol Biol Evol. 2017 Feb 1;34(2):247-261. doi: 10.1093/molbev/msw223. Mol Biol Evol. 2017. PMID: 27795231 Free PMC article.
-
Subcellular Relocalization and Positive Selection Play Key Roles in the Retention of Duplicate Genes of Populus Class III Peroxidase Family.Plant Cell. 2014 Jun;26(6):2404-2419. doi: 10.1105/tpc.114.124750. Epub 2014 Jun 16. Plant Cell. 2014. PMID: 24934172 Free PMC article.
-
Allelic variants in the PRR37 gene and the human-mediated dispersal and diversification of sorghum.Theor Appl Genet. 2015 Sep;128(9):1669-83. doi: 10.1007/s00122-015-2523-z. Epub 2015 May 16. Theor Appl Genet. 2015. PMID: 25982128
-
Plant Actin-Depolymerizing Factors Possess Opposing Biochemical Properties Arising from Key Amino Acid Changes throughout Evolution.Plant Cell. 2017 Feb;29(2):395-408. doi: 10.1105/tpc.16.00690. Epub 2017 Jan 25. Plant Cell. 2017. PMID: 28123105 Free PMC article.
-
Integrated metabolic profiling and transcriptome analysis of pigment accumulation in Lonicera japonica flower petals during colour-transition.BMC Plant Biol. 2021 Feb 17;21(1):98. doi: 10.1186/s12870-021-02877-y. BMC Plant Biol. 2021. PMID: 33596836 Free PMC article.
References
-
- Arendt J, Reznick D. Convergence and parallelism reconsidered: what have we learned about the genetics of adaptation? Trends Ecol Evol. 2008;23:26–32. - PubMed
-
- Beld M, Martin C, Huits H, Stuitje AR, Gerats AGM. Flavonoid synthesis in Petunia hybrida—partial characterization of dihydroflavonol-4-reductase genes. Plant Mol Biol. 1989;13:491–502. - PubMed
-
- Betancourt A. Lack of evidence for sign epistasis between beneficial mutations in an RNA bacteriophage. J Mol Evol. 2010;71:437–443. - PubMed
-
- Bremer B, Bremer K, Heidari N, Erixon P, Olmstead RG, Anderberg AA, Kallersjo M, Barkhordarian E. Phylogenetics of asterids based on 3 coding and 3 non-coding chloroplast DNA markers and the utility of non-coding DNA at higher taxonomic levels. Mol Phylogenet Evol. 2002;24:274–301. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

