Mos1-mediated transgenesis to probe consequences of single gene mutations in variation-rich isolates of Caenorhabditis elegans
- PMID: 23155404
- PMCID: PMC3498238
- DOI: 10.1371/journal.pone.0048762
Mos1-mediated transgenesis to probe consequences of single gene mutations in variation-rich isolates of Caenorhabditis elegans
Abstract
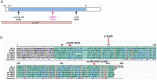
Caenorhabditis elegans, especially the N2 isolate, is an invaluable biological model system. Numerous additional natural C. elegans isolates have been shown to have unexpected genotypic and phenotypic variations which has encouraged researchers to use next generation sequencing methodology to develop a more complete picture of genotypic variations among the isolates. To understand the phenotypic effects of a genomic variation (GV) on a single gene, in a variation-rich genetic background, one should analyze that particular GV in a well understood genetic background. In C. elegans, the analysis is usually done in N2, which requires extensive crossing to bring in the GV. This can be a very time consuming procedure thus it is important to establish a fast and efficient approach to test the effect of GVs from different isolates in N2. Here we use a Mos1-mediated single-copy insertion (MosSCI) method for phenotypic assessments of GVs from the variation-rich Hawaiian strain CB4856 in N2. Specifically, we investigate effects of variations identified in the CB4856 strain on tac-1 which is an essential gene that is necessary for mitotic spindle elongation and pronuclear migration. We show the usefulness of the MosSCI method by using EU1004 tac-1(or402) as a control. or402 is a temperature sensitive lethal allele within a well-conserved TACC domain (transforming acidic coiled-coil) that results in a leucine to phenylalanine change at amino acid 229. CB4856 contains a variation that affects the second exon of tac-1 causing a cysteine to tryptophan change at amino acid 94 also within the TACC domain. Using the MosSCI method, we analyze tac-1 from CB4856 in the N2 background and demonstrate that the C94W change, albeit significant, does not cause any obvious decrease in viability. This MosSCI method has proven to be a rapid and efficient way to analyze GVs.
Conflict of interest statement
Figures

Similar articles
-
Genome-wide variations in a natural isolate of the nematode Caenorhabditis elegans.BMC Genomics. 2014 Apr 2;15:255. doi: 10.1186/1471-2164-15-255. BMC Genomics. 2014. PMID: 24694239 Free PMC article.
-
Punctuated Loci on Chromosome IV Determine Natural Variation in Orsay Virus Susceptibility of Caenorhabditis elegans Strains Bristol N2 and Hawaiian CB4856.J Virol. 2021 May 24;95(12):e02430-20. doi: 10.1128/JVI.02430-20. Print 2021 May 24. J Virol. 2021. PMID: 33827942 Free PMC article.
-
Shifting patterns of natural variation in the nuclear genome of caenorhabditis elegans.BMC Evol Biol. 2011 Jun 16;11:168. doi: 10.1186/1471-2148-11-168. BMC Evol Biol. 2011. PMID: 21679441 Free PMC article.
-
Insertional mutagenesis in C. elegans using the Drosophila transposon Mos1: a method for the rapid identification of mutated genes.Methods Mol Biol. 2006;351:59-73. doi: 10.1385/1-59745-151-7:59. Methods Mol Biol. 2006. PMID: 16988426 Review.
-
Harnessing the power of genetics: fast forward genetics in Caenorhabditis elegans.Mol Genet Genomics. 2021 Jan;296(1):1-20. doi: 10.1007/s00438-020-01721-6. Epub 2020 Sep 4. Mol Genet Genomics. 2021. PMID: 32888055 Review.
References
-
- Riddle DL, Blumenthal T, Meyer BJ, Priess JR (1997) Introduction to C. elegans. - PubMed
-
- C. elegans Sequencing Consortium (1998) Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282: 2012–2018. - PubMed
-
- Hillier LW, Marth GT, Quinlan AR, Dooling D, Fewell G, et al. (2008) Whole-genome sequencing and variant discovery in C. elegans. Nat Methods 5: 183–188. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

