Alterations in replication timing of cancer-related genes in malignant human breast cancer cells
- PMID: 23161755
- PMCID: PMC5719338
- DOI: 10.1002/jcb.24447
Alterations in replication timing of cancer-related genes in malignant human breast cancer cells
Abstract
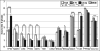
The replication timing of nine genes commonly involved in cancer was investigated in the MCF10 cell lines for human breast cancer progression. Six of these nine genes are part of a constellation of tumor suppressor genes that play a major role in familial human breast cancer (TP53, ATM, PTEN, CHK2, BRCA1, and BRCA2). Three other genes are involved in a large number of human cancers including breast as either tumor suppressors (RB1 and RAD51) or as an oncogene (cMYC). Five of these nine genes (TP53, RAD51, ATM, PTEN, and cMYC) show significant differences (P < 0.05) in replication timing between MCF10A normal human breast cells and the corresponding malignant MCF10CA1a cells. These differences are specific to the malignant state of the MCF10CA1a cells since there were no significant differences in the replication timing of these genes between normal MCF10A cells and the non-malignant cancer MCF10AT1 cells. Microarray analysis further demonstrated that three of these five genes (TP53, RAD51, and cMYC) showed significant changes in gene expression (≥2-fold) between normal and malignant cells. Our findings demonstrate an alteration in the replication timing of a small subset of cancer-related genes in malignant breast cancer cells. These alterations partially correlate with the major transcriptional changes characteristic of the malignant state in these cells.
Copyright © 2012 Wiley Periodicals, Inc.
Figures



Similar articles
-
Gene expression profiling changes induced by a novel Gemini Vitamin D derivative during the progression of breast cancer.Biochem Pharmacol. 2006 Jul 28;72(3):332-43. doi: 10.1016/j.bcp.2006.04.030. Epub 2006 May 5. Biochem Pharmacol. 2006. PMID: 16737686
-
[Promoter methylation and mRNA expression of WT1 gene in MCF10 breast cancer model].Zhonghua Bing Li Xue Za Zhi. 2007 Apr;36(4):253-8. Zhonghua Bing Li Xue Za Zhi. 2007. PMID: 17706117 Chinese.
-
Breast cancer progression in MCF10A series of cell lines is associated with alterations in retinoic acid and retinoid X receptors and with differential response to retinoids.Int J Oncol. 2004 Oct;25(4):961-71. Int J Oncol. 2004. PMID: 15375546
-
Therapeutic exploitation of tumor cell defects in homologous recombination.Anticancer Agents Med Chem. 2008 May;8(4):448-60. doi: 10.2174/187152008784220267. Anticancer Agents Med Chem. 2008. PMID: 18473729 Review.
-
Gene analysis techniques and susceptibility gene discovery in non-BRCA1/BRCA2 familial breast cancer.Surg Oncol. 2015 Jun;24(2):100-9. doi: 10.1016/j.suronc.2015.04.003. Epub 2015 Apr 13. Surg Oncol. 2015. PMID: 25936246 Review.
Cited by
-
Chromosomal Rearrangements and Altered Nuclear Organization: Recent Mechanistic Models in Cancer.Cancers (Basel). 2021 Nov 22;13(22):5860. doi: 10.3390/cancers13225860. Cancers (Basel). 2021. PMID: 34831011 Free PMC article. Review.
-
A drift-diffusion checkpoint model predicts a highly variable and growth-factor-sensitive portion of the cell cycle G1 phase.PLoS One. 2018 Feb 12;13(2):e0192087. doi: 10.1371/journal.pone.0192087. eCollection 2018. PLoS One. 2018. PMID: 29432467 Free PMC article.
-
Rapid Irreversible Transcriptional Reprogramming in Human Stem Cells Accompanied by Discordance between Replication Timing and Chromatin Compartment.Stem Cell Reports. 2019 Jul 9;13(1):193-206. doi: 10.1016/j.stemcr.2019.05.021. Epub 2019 Jun 20. Stem Cell Reports. 2019. PMID: 31231024 Free PMC article.
-
Large-scale probabilistic 3D organization of human chromosome territories.Hum Mol Genet. 2016 Feb 1;25(3):419-36. doi: 10.1093/hmg/ddv479. Epub 2015 Nov 24. Hum Mol Genet. 2016. PMID: 26604142 Free PMC article.
-
DNA replication timing, genome stability and cancer: late and/or delayed DNA replication timing is associated with increased genomic instability.Semin Cancer Biol. 2013 Apr;23(2):80-9. doi: 10.1016/j.semcancer.2013.01.001. Epub 2013 Jan 14. Semin Cancer Biol. 2013. PMID: 23327985 Free PMC article. Review.
References
-
- Amiel A, Avivi L, Gaber E, Fejgin MD. Asynchronous replication of allelic loci in Down syndrome. Eur J Hum Genet. 1998;6(4):359–64. - PubMed
-
- Amiel A, Korenstein A, Gaber E, Avivi L. Asynchronous replication of alleles in genomes carrying an extra autosome. Eur J Hum Genet. 1999;7(2):223–30. - PubMed
-
- Amiel A, Litmanovich T, Gaber E, Lishner M, Avivi L, Fejgin MD. Asynchronous replication of p53 and 21q22 loci in chronic lymphocytic leukemia. Hum Genet. 1997;101(2):219–22. - PubMed
-
- Aran D, Toperoff G, Rosenberg M, Hellman A. Replication timing-related and gene body-specific methylation of active human genes. Hum Mol Genet. 2011;20(4):670–80. - PubMed
-
- Barton MC, Crowe AJ. Chromatin alteration, transcription and replication. What’s the opening line to the story? Oncogene. 2001;20(24):3094–9. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

