Genome-wide analysis of rare copy number variations reveals PARK2 as a candidate gene for attention-deficit/hyperactivity disorder
- PMID: 23164820
- PMCID: PMC3873032
- DOI: 10.1038/mp.2012.161
Genome-wide analysis of rare copy number variations reveals PARK2 as a candidate gene for attention-deficit/hyperactivity disorder
Abstract
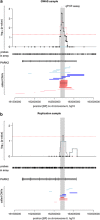
Attention-deficit/hyperactivity disorder (ADHD) is a common, highly heritable neurodevelopmental disorder. Genetic loci have not yet been identified by genome-wide association studies. Rare copy number variations (CNVs), such as chromosomal deletions or duplications, have been implicated in ADHD and other neurodevelopmental disorders. To identify rare (frequency ≤1%) CNVs that increase the risk of ADHD, we performed a whole-genome CNV analysis based on 489 young ADHD patients and 1285 adult population-based controls and identified one significantly associated CNV region. In tests for a global burden of large (>500 kb) rare CNVs, we observed a nonsignificant (P=0.271) 1.126-fold enriched rate of subjects carrying at least one such CNV in the group of ADHD cases. Locus-specific tests of association were used to assess if there were more rare CNVs in cases compared with controls. Detected CNVs, which were significantly enriched in the ADHD group, were validated by quantitative (q)PCR. Findings were replicated in an independent sample of 386 young patients with ADHD and 781 young population-based healthy controls. We identified rare CNVs within the parkinson protein 2 gene (PARK2) with a significantly higher prevalence in ADHD patients than in controls (P=2.8 × 10(-4) after empirical correction for genome-wide testing). In total, the PARK2 locus (chr 6: 162 659 756-162 767 019) harboured three deletions and nine duplications in the ADHD patients and two deletions and two duplications in the controls. By qPCR analysis, we validated 11 of the 12 CNVs in ADHD patients (P=1.2 × 10(-3) after empirical correction for genome-wide testing). In the replication sample, CNVs at the PARK2 locus were found in four additional ADHD patients and one additional control (P=4.3 × 10(-2)). Our results suggest that copy number variants at the PARK2 locus contribute to the genetic susceptibility of ADHD. Mutations and CNVs in PARK2 are known to be associated with Parkinson disease.
Figures
Similar articles
-
Rare chromosomal deletions and duplications in attention-deficit hyperactivity disorder: a genome-wide analysis.Lancet. 2010 Oct 23;376(9750):1401-8. doi: 10.1016/S0140-6736(10)61109-9. Epub 2010 Sep 29. Lancet. 2010. PMID: 20888040 Free PMC article.
-
Genome-wide copy number variation analysis in adult attention-deficit and hyperactivity disorder.J Psychiatr Res. 2014 Feb;49:60-7. doi: 10.1016/j.jpsychires.2013.10.022. Epub 2013 Nov 9. J Psychiatr Res. 2014. PMID: 24269040
-
Genome-wide analysis of copy number variants in attention deficit hyperactivity disorder: the role of rare variants and duplications at 15q13.3.Am J Psychiatry. 2012 Feb;169(2):195-204. doi: 10.1176/appi.ajp.2011.11060822. Am J Psychiatry. 2012. PMID: 22420048 Free PMC article.
-
The molecular genetic architecture of attention deficit hyperactivity disorder.Mol Psychiatry. 2015 Mar;20(3):289-97. doi: 10.1038/mp.2014.183. Epub 2015 Jan 20. Mol Psychiatry. 2015. PMID: 25600112 Review.
-
Genetics of attention-deficit/hyperactivity disorder: an update.Expert Rev Neurother. 2016;16(2):145-56. doi: 10.1586/14737175.2016.1130626. Epub 2016 Jan 11. Expert Rev Neurother. 2016. PMID: 26651394 Review.
Cited by
-
Research models of neurodevelopmental disorders: The right model in the right place.Front Neurosci. 2022 Oct 20;16:1031075. doi: 10.3389/fnins.2022.1031075. eCollection 2022. Front Neurosci. 2022. PMID: 36340790 Free PMC article. Review.
-
Genome-wide analysis of CNV (copy number variation) and their associations with narcolepsy in a Japanese population.J Hum Genet. 2014 May;59(5):235-40. doi: 10.1038/jhg.2014.13. Epub 2014 Apr 3. J Hum Genet. 2014. PMID: 24694762
-
Genome-wide analysis of copy number variations identifies PARK2 as a candidate gene for autism spectrum disorder.Mol Autism. 2016 Apr 1;7:23. doi: 10.1186/s13229-016-0087-7. eCollection 2016. Mol Autism. 2016. PMID: 27042285 Free PMC article.
-
Promising Developments in the Use of Induced Pluripotent Stem Cells in Research of ADHD.Curr Top Behav Neurosci. 2022;57:483-501. doi: 10.1007/7854_2022_346. Curr Top Behav Neurosci. 2022. PMID: 35543866
-
Energy Metabolism Disturbances in Cell Models of PARK2 CNV Carriers with ADHD.J Clin Med. 2020 Dec 18;9(12):4092. doi: 10.3390/jcm9124092. J Clin Med. 2020. PMID: 33353000 Free PMC article.
References
-
- Polanczyk G, de Lima MS, Horta BL, Biederman J, Rohde LA. The worldwide prevalence of ADHD: a systematic review and metaregression analysis. Am J Psychiatry. 2007;164:942–948. - PubMed
-
- American Psychiatric Association Diagnostic and Statistical Manual of Mental Diseases (DSM-IV)4th ed.American Psychiatric Publishing: Washington, DC; 1994
-
- Heiser P, Friedel S, Dempfle A, Konrad K, Smidt J, Grabarkiewicz J, et al. Molecular genetic aspects of attention-deficit/hyperactivity disorder. Neurosci Biobehav Rev. 2004;28:625–641. - PubMed
-
- Faraone SV, Perlis RH, Doyle AE, Smoller JW, Goralnick JJ, Holmgren MA, et al. Molecular genetics of attention deficit/hyperactivity disorder. Biol Psychiatry. 2005;57:1313–1323. - PubMed

