Lessons from model organisms: phenotypic robustness and missing heritability in complex disease
- PMID: 23166511
- PMCID: PMC3499356
- DOI: 10.1371/journal.pgen.1003041
Lessons from model organisms: phenotypic robustness and missing heritability in complex disease
Abstract
Genetically tractable model organisms from phages to mice have taught us invaluable lessons about fundamental biological processes and disease-causing mutations. Owing to technological and computational advances, human biology and the causes of human diseases have become accessible as never before. Progress in identifying genetic determinants for human diseases has been most remarkable for Mendelian traits. In contrast, identifying genetic determinants for complex diseases such as diabetes, cancer, and cardiovascular and neurological diseases has remained challenging, despite the fact that these diseases cluster in families. Hundreds of variants associated with complex diseases have been found in genome-wide association studies (GWAS), yet most of these variants explain only a modest amount of the observed heritability, a phenomenon known as "missing heritability." The missing heritability has been attributed to many factors, mainly inadequacies in genotyping and phenotyping. We argue that lessons learned about complex traits in model organisms offer an alternative explanation for missing heritability in humans. In diverse model organisms, phenotypic robustness differs among individuals, and those with decreased robustness show increased penetrance of mutations and express previously cryptic genetic variation. We propose that phenotypic robustness also differs among humans and that individuals with lower robustness will be more responsive to genetic and environmental perturbations and hence susceptible to disease. Phenotypic robustness is a quantitative trait that can be accurately measured in model organisms, but not as yet in humans. We propose feasible approaches to measure robustness in large human populations, proof-of-principle experiments for robustness markers in model organisms, and a new GWAS design that takes differences in robustness into account.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
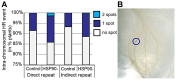
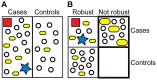
Similar articles
-
Inferring the Nature of Missing Heritability in Human Traits Using Data from the GWAS Catalog.Genetics. 2019 Jul;212(3):891-904. doi: 10.1534/genetics.119.302077. Epub 2019 May 13. Genetics. 2019. PMID: 31123044 Free PMC article.
-
The mystery of missing heritability: Genetic interactions create phantom heritability.Proc Natl Acad Sci U S A. 2012 Jan 24;109(4):1193-8. doi: 10.1073/pnas.1119675109. Epub 2012 Jan 5. Proc Natl Acad Sci U S A. 2012. PMID: 22223662 Free PMC article.
-
Underestimation of heritability using a mixed model with a polygenic covariance structure in a genome-wide association study for complex traits.Eur J Hum Genet. 2014 Jun;22(6):851-4. doi: 10.1038/ejhg.2013.236. Epub 2013 Oct 23. Eur J Hum Genet. 2014. PMID: 24149545 Free PMC article.
-
Missing heritability of complex diseases: Enlightenment by genetic variants from intermediate phenotypes.Bioessays. 2016 Jul;38(7):664-73. doi: 10.1002/bies.201600084. Epub 2016 May 31. Bioessays. 2016. PMID: 27241833 Free PMC article. Review.
-
Finding the missing heritability of complex diseases.Nature. 2009 Oct 8;461(7265):747-53. doi: 10.1038/nature08494. Nature. 2009. PMID: 19812666 Free PMC article. Review.
Cited by
-
Baldspot/ELOVL6 is a conserved modifier of disease and the ER stress response.PLoS Genet. 2018 Aug 6;14(8):e1007557. doi: 10.1371/journal.pgen.1007557. eCollection 2018 Aug. PLoS Genet. 2018. PMID: 30081392 Free PMC article.
-
A model of developmental canalization, applied to human cranial form.PLoS Comput Biol. 2021 Feb 16;17(2):e1008381. doi: 10.1371/journal.pcbi.1008381. eCollection 2021 Feb. PLoS Comput Biol. 2021. PMID: 33591964 Free PMC article.
-
Translesion Synthesis in Plants: Ultraviolet Resistance and Beyond.Front Plant Sci. 2019 Oct 9;10:1208. doi: 10.3389/fpls.2019.01208. eCollection 2019. Front Plant Sci. 2019. PMID: 31649692 Free PMC article. Review.
-
Beware of risk for increased false positive rates in genome-wide association studies for phenotypic variability.Front Genet. 2013 May 21;4:93. doi: 10.3389/fgene.2013.00093. eCollection 2013. Front Genet. 2013. PMID: 23734164 Free PMC article. No abstract available.
-
A Genome-Wide Association Analysis Reveals Epistatic Cancellation of Additive Genetic Variance for Root Length in Arabidopsis thaliana.PLoS Genet. 2015 Sep 23;11(9):e1005541. doi: 10.1371/journal.pgen.1005541. eCollection 2015. PLoS Genet. 2015. PMID: 26397943 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

