Comparative analysis of genome sequences covering the seven cronobacter species
- PMID: 23166675
- PMCID: PMC3500316
- DOI: 10.1371/journal.pone.0049455
Comparative analysis of genome sequences covering the seven cronobacter species
Abstract
Background: Species of Cronobacter are widespread in the environment and are occasional food-borne pathogens associated with serious neonatal diseases, including bacteraemia, meningitis, and necrotising enterocolitis. The genus is composed of seven species: C. sakazakii, C. malonaticus, C. turicensis, C. dublinensis, C. muytjensii, C. universalis, and C. condimenti. Clinical cases are associated with three species, C. malonaticus, C. turicensis and, in particular, with C. sakazakii multilocus sequence type 4. Thus, it is plausible that virulence determinants have evolved in certain lineages.
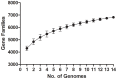
Methodology/principal findings: We generated high quality sequence drafts for eleven Cronobacter genomes representing the seven Cronobacter species, including an ST4 strain of C. sakazakii. Comparative analysis of these genomes together with the two publicly available genomes revealed Cronobacter has over 6,000 genes in one or more strains and over 2,000 genes shared by all Cronobacter. Considerable variation in the presence of traits such as type six secretion systems, metal resistance (tellurite, copper and silver), and adhesins were found. C. sakazakii is unique in the Cronobacter genus in encoding genes enabling the utilization of exogenous sialic acid which may have clinical significance. The C. sakazakii ST4 strain 701 contained additional genes as compared to other C. sakazakii but none of them were known specific virulence-related genes.
Conclusions/significance: Genome comparison revealed that pair-wise DNA sequence identity varies between 89 and 97% in the seven Cronobacter species, and also suggested various degrees of divergence. Sets of universal core genes and accessory genes unique to each strain were identified. These gene sequences can be used for designing genus/species specific detection assays. Genes encoding adhesins, T6SS, and metal resistance genes as well as prophages are found in only subsets of genomes and have contributed considerably to the variation of genomic content. Differences in gene content likely contribute to differences in the clinical and environmental distribution of species and sequence types.
Conflict of interest statement
Figures


Similar articles
-
Genome sequence of Cronobacter sakazakii BAA-894 and comparative genomic hybridization analysis with other Cronobacter species.PLoS One. 2010 Mar 8;5(3):e9556. doi: 10.1371/journal.pone.0009556. PLoS One. 2010. PMID: 20221447 Free PMC article.
-
Pan-genome analysis of the emerging foodborne pathogen Cronobacter spp. suggests a species-level bidirectional divergence driven by niche adaptation.BMC Genomics. 2013 May 31;14:366. doi: 10.1186/1471-2164-14-366. BMC Genomics. 2013. PMID: 23724777 Free PMC article.
-
Diversity of the Cronobacter genus as revealed by multilocus sequence typing.J Clin Microbiol. 2012 Sep;50(9):3031-9. doi: 10.1128/JCM.00905-12. Epub 2012 Jul 11. J Clin Microbiol. 2012. PMID: 22785185 Free PMC article.
-
Cronobacter spp.--opportunistic food-borne pathogens. A review of their virulence and environmental-adaptive traits.J Med Microbiol. 2014 Aug;63(Pt 8):1023-1037. doi: 10.1099/jmm.0.073742-0. Epub 2014 May 30. J Med Microbiol. 2014. PMID: 24878566 Review.
-
Strategies for the identification and tracking of cronobacter species: an opportunistic pathogen of concern to neonatal health.Front Pediatr. 2015 May 5;3:38. doi: 10.3389/fped.2015.00038. eCollection 2015. Front Pediatr. 2015. PMID: 26000266 Free PMC article. Review.
Cited by
-
Genome Sequence of Hydrocarbon-Degrading Cronobacter sp. Strain DJ34 Isolated from Crude Oil-Containing Sludge from the Duliajan Oil Fields, Assam, India.Genome Announc. 2015 Nov 12;3(6):e01321-15. doi: 10.1128/genomeA.01321-15. Genome Announc. 2015. PMID: 26564043 Free PMC article.
-
L-Fucose is involved in human-gut microbiome interactions.Appl Microbiol Biotechnol. 2023 Jun;107(12):3869-3875. doi: 10.1007/s00253-023-12527-y. Epub 2023 May 6. Appl Microbiol Biotechnol. 2023. PMID: 37148338 Review.
-
Host Sialic Acids: A Delicacy for the Pathogen with Discerning Taste.Microbiol Spectr. 2015 Aug;3(4):10.1128/microbiolspec.MBP-0005-2014. doi: 10.1128/microbiolspec.MBP-0005-2014. Microbiol Spectr. 2015. PMID: 26350327 Free PMC article. Review.
-
Fatal Necrotizing Enterocolitis in Neonate Caused by Cronobacter sakazakii Sequence Type 64 Strain of CRISPR Sublineage b.Emerg Infect Dis. 2023 Sep;29(9):1917-1920. doi: 10.3201/eid2909.230537. Emerg Infect Dis. 2023. PMID: 37610257 Free PMC article.
-
Complete Genome Sequence of Cronobacter dublinensis Bacteriophage vB_Cdu_VP8.Microbiol Resour Announc. 2023 Jun 20;12(6):e0029623. doi: 10.1128/mra.00296-23. Epub 2023 May 17. Microbiol Resour Announc. 2023. PMID: 37195201 Free PMC article.
References
-
- Friedemann M (2007) Enterobacter sakazakii in food and beverages (other than infant formula and milk powder). Int J Food Microbiol 116: 1–10. - PubMed
-
- Kandhai MC, Reij MW, Gorris LG, Guillaume-Gentil O, van Schothorst M (2004) Occurrence of Enterobacter sakazakii in food production environments and households. Lancet 363: 39–40. - PubMed
-
- Kucerova E, Joseph S, Forsythe SJ (2011) The Cronobacter genus: ubiquity and diversity. Quality Assurance and Safety of Crops & Foods 3: 104–122.
-
- FAO/WHO (2008) Enterobacter sakazakii (Cronobacter spp.) in powdered follow-up formulae. Microbiological Risk Assessment Series No. 15, Rome, 90pp. Available: http://www.who.int/foodsafety/publications/micro/mra_followup/en/.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

