Distinct disulfide isomers of μ-conotoxins KIIIA and KIIIB block voltage-gated sodium channels
- PMID: 23167564
- PMCID: PMC4131687
- DOI: 10.1021/bi301256s
Distinct disulfide isomers of μ-conotoxins KIIIA and KIIIB block voltage-gated sodium channels
Abstract
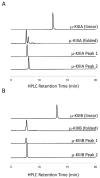
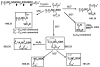
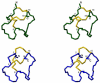
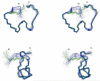
In the preparation of synthetic conotoxins containing multiple disulfide bonds, oxidative folding can produce numerous permutations of disulfide bond connectivities. Establishing the native disulfide connectivities thus presents a significant challenge when the venom-derived peptide is not available, as is increasingly the case when conotoxins are identified from cDNA sequences. Here, we investigate the disulfide connectivity of μ-conotoxin KIIIA, which was predicted originally to have a [C1-C9,C2-C15,C4-C16] disulfide pattern based on homology with closely related μ-conotoxins. The two major isomers of synthetic μ-KIIIA formed during oxidative folding were purified and their disulfide connectivities mapped by direct mass spectrometric collision-induced dissociation fragmentation of the disulfide-bonded polypeptides. Our results show that the major oxidative folding product adopts a [C1-C15,C2-C9,C4-C16] disulfide connectivity, while the minor product adopts a [C1-C16,C2-C9,C4-C15] connectivity. Both of these peptides were potent blockers of Na(V)1.2 (K(d) values of 5 and 230 nM, respectively). The solution structure for μ-KIIIA based on nuclear magnetic resonance data was recalculated with the [C1-C15,C2-C9,C4-C16] disulfide pattern; its structure was very similar to the μ-KIIIA structure calculated with the incorrect [C1-C9,C2-C15,C4-C16] disulfide pattern, with an α-helix spanning residues 7-12. In addition, the major folding isomers of μ-KIIIB, an N-terminally extended isoform of μ-KIIIA identified from its cDNA sequence, were isolated. These folding products had the same disulfide connectivities as μ-KIIIA, and both blocked Na(V)1.2 (K(d) values of 470 and 26 nM, respectively). Our results establish that the preferred disulfide pattern of synthetic μ-KIIIA and μ-KIIIB folded in vitro is 1-5/2-4/3-6 but that other disulfide isomers are also potent sodium channel blockers. These findings raise questions about the disulfide pattern(s) of μ-KIIIA in the venom of Conus kinoshitai; indeed, the presence of multiple disulfide isomers in the venom could provide a means of further expanding the snail's repertoire of active peptides.
Figures


Similar articles
-
Synthesis of the Most Potent Isomer of μ-Conotoxin KIIIA Using Different Strategies.Molecules. 2023 Apr 11;28(8):3377. doi: 10.3390/molecules28083377. Molecules. 2023. PMID: 37110612 Free PMC article.
-
Structural and functional insights into the inhibition of human voltage-gated sodium channels by μ-conotoxin KIIIA disulfide isomers.J Biol Chem. 2022 Mar;298(3):101728. doi: 10.1016/j.jbc.2022.101728. Epub 2022 Feb 12. J Biol Chem. 2022. PMID: 35167877 Free PMC article.
-
Structure of the analgesic mu-conotoxin KIIIA and effects on the structure and function of disulfide deletion.Biochemistry. 2009 Feb 17;48(6):1210-9. doi: 10.1021/bi801998a. Biochemistry. 2009. PMID: 19170536 Free PMC article.
-
Using the deadly mu-conotoxins as probes of voltage-gated sodium channels.Toxicon. 2004 Aug;44(2):117-22. doi: 10.1016/j.toxicon.2004.03.028. Toxicon. 2004. PMID: 15246758 Free PMC article. Review.
-
Folding of conotoxins: formation of the native disulfide bridges during chemical synthesis and biosynthesis of Conus peptides.Antioxid Redox Signal. 2008 Jan;10(1):141-55. doi: 10.1089/ars.2007.1856. Antioxid Redox Signal. 2008. PMID: 17961068 Review.
Cited by
-
Expanding chemical diversity of conotoxins: peptoid-peptide chimeras of the sodium channel blocker μ-KIIIA and its selenopeptide analogues.Eur J Med Chem. 2013 Jul;65:144-50. doi: 10.1016/j.ejmech.2013.04.041. Epub 2013 May 1. Eur J Med Chem. 2013. PMID: 23707919 Free PMC article.
-
Sodium channels and pain: from toxins to therapies.Br J Pharmacol. 2018 Jun;175(12):2138-2157. doi: 10.1111/bph.13962. Epub 2017 Sep 2. Br J Pharmacol. 2018. PMID: 28749537 Free PMC article. Review.
-
Total synthesis of μ-conotoxin lt5d.RSC Adv. 2018 Oct 30;8(64):36579-36583. doi: 10.1039/c8ra03706j. eCollection 2018 Oct 26. RSC Adv. 2018. PMID: 35558937 Free PMC article.
-
Predicting Structural Details of the Sodium Channel Pore Basing on Animal Toxin Studies.Front Pharmacol. 2018 Aug 7;9:880. doi: 10.3389/fphar.2018.00880. eCollection 2018. Front Pharmacol. 2018. PMID: 30131702 Free PMC article. Review.
-
Structural and Functional Analyses of Cone Snail Toxins.Mar Drugs. 2019 Jun 21;17(6):370. doi: 10.3390/md17060370. Mar Drugs. 2019. PMID: 31234371 Free PMC article. Review.
References
-
- Khoo KK, Norton RS. Role of disulfide bonds in peptide and protein conformation. In: Hughes AB, editor. Amino Acids, Peptides and Proteins in Organic Chemistry. Wiley-VCH, Weinheim; Germany: 2012. pp. 395–417.
-
- Gilbert HF. Thiol/disulfide exchange equilibria and disulfide bond stability. Methods Enzymol. 1995;251:8–28. - PubMed
-
- Holmgren A, Bjornstedt M. Thioredoxin and thioredoxin reductase. Methods Enzymol. 1995;252:199–208. - PubMed
-
- Buczek O, Green BR, Bulaj G. Albumin is a redox-active crowding agent that promotes oxidative folding of cysteine-rich peptides. Biopolymers. 2007;88:8–19. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

