Dual-acting riboswitch control of translation initiation and mRNA decay
- PMID: 23169642
- PMCID: PMC3528543
- DOI: 10.1073/pnas.1214024109
Dual-acting riboswitch control of translation initiation and mRNA decay
Abstract
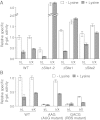
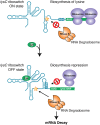
Riboswitches are mRNA regulatory elements that control gene expression by altering their structure in response to specific metabolite binding. In bacteria, riboswitches consist of an aptamer that performs ligand recognition and an expression platform that regulates either transcription termination or translation initiation. Here, we describe a dual-acting riboswitch from Escherichia coli that, in addition to modulating translation initiation, also is directly involved in the control of initial mRNA decay. Upon lysine binding, the lysC riboswitch adopts a conformation that not only inhibits translation initiation but also exposes RNase E cleavage sites located in the riboswitch expression platform. However, in the absence of lysine, the riboswitch folds into an alternative conformation that simultaneously allows translation initiation and sequesters RNase E cleavage sites. Both regulatory activities can be individually inhibited, indicating that translation initiation and mRNA decay can be modulated independently using the same conformational switch. Because RNase E cleavage sites are located in the riboswitch sequence, this riboswitch provides a unique means for the riboswitch to modulate RNase E cleavage activity directly as a function of lysine. This dual inhibition is in contrast to other riboswitches, such as the thiamin pyrophosphate-sensing thiM riboswitch, which triggers mRNA decay only as a consequence of translation inhibition. The riboswitch control of RNase E cleavage activity is an example of a mechanism by which metabolite sensing is used to regulate gene expression of single genes or even large polycistronic mRNAs as a function of environmental changes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

