The interactions of GW182 proteins with PABP and deadenylases are required for both translational repression and degradation of miRNA targets
- PMID: 23172285
- PMCID: PMC3553986
- DOI: 10.1093/nar/gks1078
The interactions of GW182 proteins with PABP and deadenylases are required for both translational repression and degradation of miRNA targets
Abstract
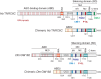
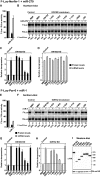
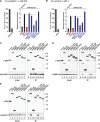
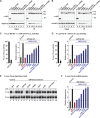
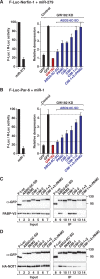
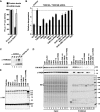
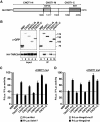
Animal miRNAs silence the expression of mRNA targets through translational repression, deadenylation and subsequent mRNA degradation. Silencing requires association of miRNAs with an Argonaute protein and a GW182 family protein. In turn, GW182 proteins interact with poly(A)-binding protein (PABP) and the PAN2-PAN3 and CCR4-NOT deadenylase complexes. These interactions are required for the deadenylation and decay of miRNA targets. Recent studies have indicated that miRNAs repress translation before inducing target deadenylation and decay; however, whether translational repression and deadenylation are coupled or represent independent repressive mechanisms is unclear. Another remaining question is whether translational repression also requires GW182 proteins to interact with both PABP and deadenylases. To address these questions, we characterized the interaction of Drosophila melanogaster GW182 with deadenylases and defined the minimal requirements for a functional GW182 protein. Functional assays in D. melanogaster and human cells indicate that miRNA-mediated translational repression and degradation are mechanistically linked and are triggered through the interactions of GW182 proteins with PABP and deadenylases.
Figures


Similar articles
-
GW182 proteins cause PABP dissociation from silenced miRNA targets in the absence of deadenylation.EMBO J. 2013 Apr 3;32(7):1052-65. doi: 10.1038/emboj.2013.44. Epub 2013 Mar 5. EMBO J. 2013. PMID: 23463101 Free PMC article.
-
GW182 proteins directly recruit cytoplasmic deadenylase complexes to miRNA targets.Mol Cell. 2011 Oct 7;44(1):120-33. doi: 10.1016/j.molcel.2011.09.007. Mol Cell. 2011. PMID: 21981923
-
The role of GW182 proteins in miRNA-mediated gene silencing.Adv Exp Med Biol. 2013;768:147-63. doi: 10.1007/978-1-4614-5107-5_9. Adv Exp Med Biol. 2013. PMID: 23224969 Review.
-
miRNA-mediated deadenylation is orchestrated by GW182 through two conserved motifs that interact with CCR4-NOT.Nat Struct Mol Biol. 2011 Oct 7;18(11):1211-7. doi: 10.1038/nsmb.2149. Nat Struct Mol Biol. 2011. PMID: 21984185
-
A molecular link between miRISCs and deadenylases provides new insight into the mechanism of gene silencing by microRNAs.Cold Spring Harb Perspect Biol. 2012 Dec 1;4(12):a012328. doi: 10.1101/cshperspect.a012328. Cold Spring Harb Perspect Biol. 2012. PMID: 23209154 Free PMC article. Review.
Cited by
-
Coordinated Regulation of Cap-Dependent Translation and MicroRNA Function by Convergent Signaling Pathways.Mol Cell Biol. 2016 Aug 26;36(18):2360-73. doi: 10.1128/MCB.01011-15. Print 2016 Sep 15. Mol Cell Biol. 2016. PMID: 27354062 Free PMC article.
-
Ribosome dynamics and mRNA turnover, a complex relationship under constant cellular scrutiny.Wiley Interdiscip Rev RNA. 2021 Nov;12(6):e1658. doi: 10.1002/wrna.1658. Epub 2021 May 5. Wiley Interdiscip Rev RNA. 2021. PMID: 33949788 Free PMC article. Review.
-
Postnatal liver functional maturation requires Cnot complex-mediated decay of mRNAs encoding cell cycle and immature liver genes.Development. 2019 Feb 15;146(4):dev168146. doi: 10.1242/dev.168146. Development. 2019. PMID: 30733279 Free PMC article.
-
Translation Initiation Regulated by RNA-Binding Protein in Mammals: The Modulation of Translation Initiation Complex by Trans-Acting Factors.Cells. 2021 Jul 6;10(7):1711. doi: 10.3390/cells10071711. Cells. 2021. PMID: 34359885 Free PMC article. Review.
-
A Compilation of the Diverse miRNA Functions in Caenorhabditis elegans and Drosophila melanogaster Development.Int J Mol Sci. 2023 Apr 9;24(8):6963. doi: 10.3390/ijms24086963. Int J Mol Sci. 2023. PMID: 37108126 Free PMC article. Review.
References
-
- Huntzinger E, Izaurralde E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat. Rev. Genet. 2011;12:99–110. - PubMed
-
- Fabian MR, Sonenberg N. The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC. Nat. Struct. Mol. Biol. 2012;19:586–593. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

