On the role of PDZ domain-encoding genes in Drosophila border cell migration
- PMID: 23173089
- PMCID: PMC3484668
- DOI: 10.1534/g3.112.004093
On the role of PDZ domain-encoding genes in Drosophila border cell migration
Abstract
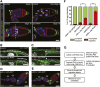
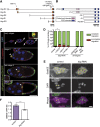
Cells often move as collective groups during normal embryonic development and wound healing, although the mechanisms governing this type of migration are poorly understood. The Drosophila melanogaster border cells migrate as a cluster during late oogenesis and serve as a powerful in vivo genetic model for collective cell migration. To discover new genes that participate in border cell migration, 64 out of 66 genes that encode PDZ domain-containing proteins were systematically targeted by in vivo RNAi knockdown. The PDZ domain is one of the largest families of protein-protein interaction domains found in eukaryotes. Proteins that contain PDZ domains participate in a variety of biological processes, including signal transduction and establishment of epithelial apical-basal polarity. Targeting PDZ proteins effectively assesses a larger number of genes via the protein complexes and pathways through which these proteins function. par-6, a known regulator of border cell migration, was a positive hit and thus validated the approach. Knockdown of 14 PDZ domain genes disrupted migration with multiple RNAi lines. The candidate genes have diverse predicted cellular functions and are anticipated to provide new insights into the mechanisms that control border cell movement. As a test of this concept, two genes that disrupted migration were characterized in more detail: big bang and the Dlg5 homolog CG6509. We present evidence that Big bang regulates JAK/STAT signaling, whereas Dlg5/CG6509 maintains cluster cohesion. Moreover, these results demonstrate that targeting a selected class of genes by RNAi can uncover novel regulators of collective cell migration.
Keywords: Drosophila; JAK/STAT; PSD95/Dlg/ZO-1 (PDZ) domains; border cells; collective migration.
Figures



References
-
- Abdelilah-Seyfried S., Cox D. N., Jan Y. N., 2003. Bazooka is a permissive factor for the invasive behavior of discs large tumor cells in Drosophila ovarian follicular epithelia. Development 130: 1927–1935 - PubMed
-
- Bai J., Uehara Y., Montell D. J., 2000. Regulation of invasive cell behavior by taiman, a Drosophila protein related to AIB1, a steroid receptor coactivator amplified in breast cancer. Cell 103: 1047–1058 - PubMed
-
- Beccari S., Teixeira L., Rørth P., 2002. The JAK/STAT pathway is required for border cell migration during Drosophila oogenesis. Mech. Dev. 111: 115–123 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
