Function and regulation of SUMO proteases
- PMID: 23175280
- PMCID: PMC3668692
- DOI: 10.1038/nrm3478
Function and regulation of SUMO proteases
Abstract
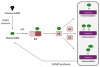
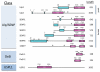
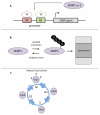
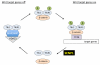
Covalent attachment of small ubiquitin-like modifier (SUMO) to proteins is highly dynamic, and both SUMO-protein conjugation and cleavage can be regulated. Protein desumoylation is carried out by SUMO proteases, which control cellular mechanisms ranging from transcription and cell division to ribosome biogenesis. Recent advances include the discovery of two novel classes of SUMO proteases, insights regarding SUMO protease specificity, and revelations of previously unappreciated SUMO protease functions in several key cellular pathways. These developments, together with new connections between SUMO proteases and the recently discovered SUMO-targeted ubiquitin ligases (STUbLs), make this an exciting period to study these enzymes.
Figures


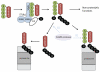
References
-
- Panse VG, Hardeland U, Werner T, Kuster B, Hurt E. A proteome-wide approach identifies sumoylated substrate proteins in yeast. J Biol Chem. 2004;279:41346–51. - PubMed
-
- Wohlschlegel JA, Johnson ES, Reed SI, Yates JR., 3rd Global analysis of protein sumoylation in Saccharomyces cerevisiae. J Biol Chem. 2004;279:45662–8. - PubMed
-
- Hannich JT, et al. Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae. J Biol Chem. 2005;280:4102–10. - PubMed
-
- Wykoff DD, O'Shea EK. Identification of sumoylated proteins by systematic immunoprecipitation of the budding yeast proteome. Mol Cell Proteomics. 2005;4:73–83. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

