Position-dependent splicing activation and repression by SR and hnRNP proteins rely on common mechanisms
- PMID: 23175589
- PMCID: PMC3527730
- DOI: 10.1261/rna.037044.112
Position-dependent splicing activation and repression by SR and hnRNP proteins rely on common mechanisms
Erratum in
- RNA. 2013 Jul;19(7):1015
Abstract
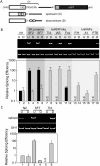
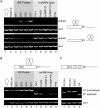
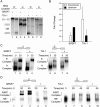
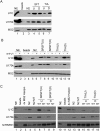
Alternative splicing is regulated by splicing factors that modulate splice site selection. In some cases, however, splicing factors show antagonistic activities by either activating or repressing splicing. Here, we show that these opposing outcomes are based on their binding location relative to regulated 5' splice sites. SR proteins enhance splicing only when they are recruited to the exon. However, they interfere with splicing by simply relocating them to the opposite intronic side of the splice site. hnRNP splicing factors display analogous opposing activities, but in a reversed position dependence. Activation by SR or hnRNP proteins increases splice site recognition at the earliest steps of exon definition, whereas splicing repression promotes the assembly of nonproductive complexes that arrest spliceosome assembly prior to splice site pairing. Thus, SR and hnRNP splicing factors exploit similar mechanisms to positively or negatively influence splice site selection.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
