Kinetic competition between RNA Polymerase II and Sen1-dependent transcription termination
- PMID: 23177741
- PMCID: PMC3545030
- DOI: 10.1016/j.molcel.2012.10.014
Kinetic competition between RNA Polymerase II and Sen1-dependent transcription termination
Abstract
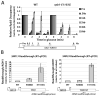
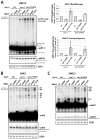
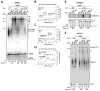
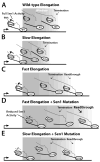
The essential helicase-like protein Sen1 mediates termination of RNA Polymerase II (Pol II) transcription at snoRNAs and other noncoding RNAs in yeast. A mutation in the Pol II subunit Rpb1 that increases the elongation rate increases read-through transcription at Sen1-mediated terminators. Termination and growth defects in sen1 mutant cells are partially suppressed by a slowly transcribing Pol II mutant and are exacerbated by a faster-transcribing Pol II mutant. Deletion of the nuclear exosome subunit Rrp6 allows visualization of noncoding RNA intermediates that are terminated but not yet processed. Sen1 mutants or faster-transcribing Pol II increase the average lengths of preprocessed snoRNA, CUT, and SUT transcripts, while slowed Pol II transcription produces shorter transcripts. These connections between transcription rate and Sen1 activity support a model whereby kinetic competition between elongating Pol II and Sen1 helicase establishes the temporal and spatial window for early Pol II termination.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Biochemical characterization of the helicase Sen1 provides new insights into the mechanisms of non-coding transcription termination.Nucleic Acids Res. 2017 Feb 17;45(3):1355-1370. doi: 10.1093/nar/gkw1230. Nucleic Acids Res. 2017. PMID: 28180347 Free PMC article.
-
RNA Polymerase II Transcription Attenuation at the Yeast DNA Repair Gene, DEF1, Involves Sen1-Dependent and Polyadenylation Site-Dependent Termination.G3 (Bethesda). 2018 May 31;8(6):2043-2058. doi: 10.1534/g3.118.200072. G3 (Bethesda). 2018. PMID: 29686108 Free PMC article.
-
Single-molecule characterization of extrinsic transcription termination by Sen1 helicase.Nat Commun. 2019 Apr 4;10(1):1545. doi: 10.1038/s41467-019-09560-9. Nat Commun. 2019. PMID: 30948716 Free PMC article.
-
The Nrd1-Nab3-Sen1 transcription termination complex from a structural perspective.Biochem Soc Trans. 2023 Jun 28;51(3):1257-1269. doi: 10.1042/BST20221418. Biochem Soc Trans. 2023. PMID: 37222282 Free PMC article. Review.
-
Transcription termination and the control of the transcriptome: why, where and how to stop.Nat Rev Mol Cell Biol. 2015 Mar;16(3):190-202. doi: 10.1038/nrm3943. Epub 2015 Feb 4. Nat Rev Mol Cell Biol. 2015. PMID: 25650800 Review.
Cited by
-
RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity.Cell. 2015 May 7;161(4):774-89. doi: 10.1016/j.cell.2015.04.034. Cell. 2015. PMID: 25957685 Free PMC article.
-
Termination of non-coding transcription in yeast relies on both an RNA Pol II CTD interaction domain and a CTD-mimicking region in Sen1.EMBO J. 2020 Apr 1;39(7):e101548. doi: 10.15252/embj.2019101548. Epub 2020 Feb 28. EMBO J. 2020. PMID: 32107786 Free PMC article.
-
Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons.Elife. 2014 Apr 29;3:e02407. doi: 10.7554/eLife.02407. Elife. 2014. PMID: 24843027 Free PMC article.
-
KAP1 negatively regulates RNA polymerase II elongation kinetics to activate signal-induced transcription.bioRxiv [Preprint]. 2024 May 5:2024.05.05.592422. doi: 10.1101/2024.05.05.592422. bioRxiv. 2024. Update in: Nat Commun. 2024 Jul 12;15(1):5859. doi: 10.1038/s41467-024-49905-7. PMID: 38746145 Free PMC article. Updated. Preprint.
-
Single-molecule reconstruction of eukaryotic factor-dependent transcription termination.Nat Commun. 2024 Jun 15;15(1):5113. doi: 10.1038/s41467-024-49527-z. Nat Commun. 2024. PMID: 38879529 Free PMC article.
References
-
- Arigo JT, Carroll KL, Ames JM, Corden JL. Regulation of yeast NRD1 expression by premature transcription termination. Mol Cell. 2006;21:641–651. - PubMed
-
- Bar-Nahum G, Epshtein V, Ruckenstein AE, Rafikov R, Mustaev A, Nudler E. A ratchet mechanism of transcription elongation and its control. Cell. 2005;120:183–193. - PubMed
-
- Bleichert F, Baserga SJ. The long unwinding road of RNA helicases. Mol Cell. 2007;27:339–352. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

