AID and Apobec3G haphazard deamination and mutational diversity
- PMID: 23178850
- PMCID: PMC3600397
- DOI: 10.1007/s00018-012-1212-1
AID and Apobec3G haphazard deamination and mutational diversity
Abstract
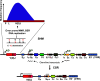
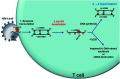
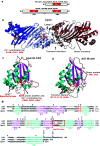
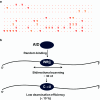
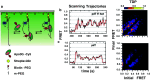
Activation-induced deoxycytidine deaminase (AID) and Apobec 3G (Apo3G) cause mutational diversity by initiating mutations on regions of single-stranded (ss) DNA. Expressed in B cells, AID deaminates C → U in actively transcribed immunoglobulin (Ig) variable and switch regions to initiate the somatic hypermutation (SHM) and class switch recombination (CSR) that are essential for antibody diversity. Apo3G expressed in T cells catalyzes C deaminations on reverse transcribed cDNA causing HIV-1 retroviral inactivation. When operating properly, AID- and Apo3G-initiated mutations boost human fitness. Yet, both enzymes are potentially powerful somatic cell "mutators". Loss of regulated expression and proper genome targeting can cause human cancer. Here, we review well-established biological roles of AID and Apo3G. We provide a synopsis of AID partnering proteins during SHM and CSR, and describe how an Apo2 crystal structure provides "surrogate" insight for AID and Apo3G biochemical behavior. However, large gaps remain in our understanding of how dC deaminases search ssDNA to identify trinucleotide motifs to deaminate. We discuss two recent methods to analyze ssDNA scanning and deamination. Apo3G scanning and deamination is visualized in real-time using single-molecule FRET, and AID deamination efficiencies are determined with a random walk analysis. AID and Apo3G encounter many candidate deamination sites while scanning ssDNA. Generating mutational diversity is a principal aim of AID and an important ancillary property of Apo3G. Success seems likely to involve hit and miss deamination motif targeting, biased strongly toward miss.
Figures

Similar articles
-
Single-stranded DNA scanning and deamination by APOBEC3G cytidine deaminase at single molecule resolution.J Biol Chem. 2012 May 4;287(19):15826-35. doi: 10.1074/jbc.M112.342790. Epub 2012 Feb 23. J Biol Chem. 2012. PMID: 22362763 Free PMC article.
-
Intensity of deoxycytidine deamination of HIV-1 proviral DNA by the retroviral restriction factor APOBEC3G is mediated by the noncatalytic domain.J Biol Chem. 2011 Apr 1;286(13):11415-26. doi: 10.1074/jbc.M110.199604. Epub 2011 Feb 7. J Biol Chem. 2011. PMID: 21300806 Free PMC article.
-
Biochemical analysis of hypermutation by the deoxycytidine deaminase APOBEC3A.J Biol Chem. 2012 Aug 31;287(36):30812-22. doi: 10.1074/jbc.M112.393181. Epub 2012 Jul 20. J Biol Chem. 2012. PMID: 22822074 Free PMC article.
-
Stochastic properties of processive cytidine DNA deaminases AID and APOBEC3G.Philos Trans R Soc Lond B Biol Sci. 2009 Mar 12;364(1517):583-93. doi: 10.1098/rstb.2008.0195. Philos Trans R Soc Lond B Biol Sci. 2009. PMID: 19022738 Free PMC article. Review.
-
AID: how does it aid antibody diversity?Immunity. 2004 Jun;20(6):659-68. doi: 10.1016/j.immuni.2004.05.011. Immunity. 2004. PMID: 15189732 Review.
Cited by
-
Tautomerism provides a molecular explanation for the mutagenic properties of the anti-HIV nucleoside 5-aza-5,6-dihydro-2'-deoxycytidine.Proc Natl Acad Sci U S A. 2014 Aug 12;111(32):E3252-9. doi: 10.1073/pnas.1405635111. Epub 2014 Jul 28. Proc Natl Acad Sci U S A. 2014. PMID: 25071207 Free PMC article.
-
Molecular targets and pathways involved in liver metastasis of colorectal cancer.Clin Exp Metastasis. 2015 Aug;32(6):623-35. doi: 10.1007/s10585-015-9732-3. Epub 2015 Jun 24. Clin Exp Metastasis. 2015. PMID: 26104118 Review.
-
Random Walk Enzymes: Information Theory, Quantum Isomorphism, and Entropy Dispersion.J Phys Chem A. 2019 Apr 4;123(13):3030-3037. doi: 10.1021/acs.jpca.9b00910. Epub 2019 Mar 21. J Phys Chem A. 2019. PMID: 30848911 Free PMC article.
-
Mutations in human AID differentially affect its ability to deaminate cytidine and 5-methylcytidine in ssDNA substrates in vitro.Sci Rep. 2017 Jun 20;7(1):3873. doi: 10.1038/s41598-017-03936-x. Sci Rep. 2017. PMID: 28634398 Free PMC article.
-
The multifaceted roles of RNA binding in APOBEC cytidine deaminase functions.Wiley Interdiscip Rev RNA. 2014 Jul-Aug;5(4):493-508. doi: 10.1002/wrna.1226. Epub 2014 Mar 24. Wiley Interdiscip Rev RNA. 2014. PMID: 24664896 Free PMC article. Review.
References
-
- Milstein C. Diversity and the genesis of high affinity antibodies. Biochem Soc Trans. 1987;15:779–787. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

