miR-200c targets a NF-κB up-regulated TrkB/NTF3 autocrine signaling loop to enhance anoikis sensitivity in triple negative breast cancer
- PMID: 23185507
- PMCID: PMC3503774
- DOI: 10.1371/journal.pone.0049987
miR-200c targets a NF-κB up-regulated TrkB/NTF3 autocrine signaling loop to enhance anoikis sensitivity in triple negative breast cancer
Abstract
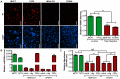
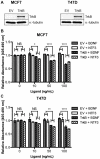
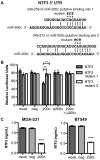
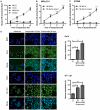
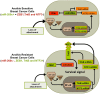
Anoikis is apoptosis initiated upon cell detachment from the native extracellular matrix. Since survival upon detachment from basement membrane is required for metastasis, the ability to resist anoikis contributes to the metastatic potential of breast tumors. miR-200c, a potent repressor of epithelial to mesenchymal transition, is expressed in luminal breast cancers, but is lost in more aggressive basal-like, or triple negative breast cancers (TNBC). We previously demonstrated that miR-200c restores anoikis sensitivity to TNBC cells by directly targeting the neurotrophic receptor tyrosine kinase, TrkB. In this study, we identify a TrkB ligand, neurotrophin 3 (NTF3), as capable of activating TrkB to induce anoikis resistance, and show that NTF3 is also a direct target of miR-200c. We present the first evidence that anoikis resistant TNBC cells up-regulate both TrkB and NTF3 when suspended, and show that this up-regulation is necessary for survival in suspension. We further demonstrate that NF-κB activity increases 6 fold in suspended TNBC cells, and identify RelA and NF-κB1 as the transcription factors responsible for suspension-induced up-regulation of TrkB and NTF3. Consequently, inhibition of NF-κB activity represses anoikis resistance. Taken together, our findings define a critical mechanism for transcriptional and post-transcriptional control of suspension-induced up-regulation of TrkB and NTF3 in anoikis resistant breast cancer cells.
Conflict of interest statement
Figures



Similar articles
-
Targets of miR-200c mediate suppression of cell motility and anoikis resistance.Breast Cancer Res. 2011 Apr 18;13(2):R45. doi: 10.1186/bcr2867. Breast Cancer Res. 2011. PMID: 21501518 Free PMC article.
-
Upregulation of TrkB promotes epithelial-mesenchymal transition and anoikis resistance in endometrial carcinoma.PLoS One. 2013 Jul 30;8(7):e70616. doi: 10.1371/journal.pone.0070616. Print 2013. PLoS One. 2013. PMID: 23936232 Free PMC article.
-
A TDO2-AhR signaling axis facilitates anoikis resistance and metastasis in triple-negative breast cancer.Cancer Res. 2015 Nov 1;75(21):4651-64. doi: 10.1158/0008-5472.CAN-15-2011. Epub 2015 Sep 11. Cancer Res. 2015. PMID: 26363006 Free PMC article.
-
Anoikis molecular pathways and its role in cancer progression.Biochim Biophys Acta. 2013 Dec;1833(12):3481-3498. doi: 10.1016/j.bbamcr.2013.06.026. Epub 2013 Jul 2. Biochim Biophys Acta. 2013. PMID: 23830918 Review.
-
Regulators and mechanisms of anoikis in triple-negative breast cancer (TNBC): A review.Crit Rev Oncol Hematol. 2019 Aug;140:17-27. doi: 10.1016/j.critrevonc.2019.05.009. Epub 2019 May 18. Crit Rev Oncol Hematol. 2019. PMID: 31154235 Review.
Cited by
-
Friend or foe: emerging role of nuclear factor kappa-light-chain-enhancer of activated B cells in cell senescence.Onco Targets Ther. 2013 Sep 4;6:1221-9. doi: 10.2147/OTT.S36160. Onco Targets Ther. 2013. PMID: 24043947 Free PMC article. Review.
-
An integrated analysis of the SOX2 microRNA response program in human pluripotent and nullipotent stem cell lines.BMC Genomics. 2014 Aug 25;15:711. doi: 10.1186/1471-2164-15-711. BMC Genomics. 2014. PMID: 25156079 Free PMC article.
-
Targeting the NFκB signaling pathways for breast cancer prevention and therapy.Curr Med Chem. 2015;22(2):264-89. doi: 10.2174/0929867321666141106124315. Curr Med Chem. 2015. PMID: 25386819 Free PMC article. Review.
-
Silencing LRH-1 in colon cancer cell lines impairs proliferation and alters gene expression programs.Proc Natl Acad Sci U S A. 2015 Feb 24;112(8):2467-72. doi: 10.1073/pnas.1500978112. Epub 2015 Feb 9. Proc Natl Acad Sci U S A. 2015. PMID: 25675535 Free PMC article.
-
Six immune-related promising biomarkers may promote hepatocellular carcinoma prognosis: a bioinformatics analysis and experimental validation.Cancer Cell Int. 2023 Mar 23;23(1):52. doi: 10.1186/s12935-023-02888-9. Cancer Cell Int. 2023. PMID: 36959615 Free PMC article.
References
-
- Siegel R, Naishadham D, Jemal A (2012) Cancer statistics, 2012. CA Cancer J Clin 62: 10–29 doi:10.3322/caac.20138 - DOI - PubMed
-
- Weigelt B, Peterse JL, van ’t Veer LJ (2005) Breast cancer metastasis: markers and models. Nat Rev Cancer 5: 591–602 doi:10.1038/nrc1670 - DOI - PubMed
-
- Drasin DJ, Robin TP, Ford HL (2011) Breast cancer epithelial-to-mesenchymal transition: examining the functional consequences of plasticity. Breast Cancer Res 13: 226 doi:10.1186/bcr3037 - DOI - PMC - PubMed
-
- Pillai RS, Bhattacharyya SN, Filipowicz W (2007) Repression of protein synthesis by miRNAs: how many mechanisms? Trends in Cell Biology 17: 118–126. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

