Pyrimidine salvage in Trypanosoma brucei bloodstream forms and the trypanocidal action of halogenated pyrimidines
- PMID: 23188714
- PMCID: PMC4857052
- DOI: 10.1124/mol.112.082321
Pyrimidine salvage in Trypanosoma brucei bloodstream forms and the trypanocidal action of halogenated pyrimidines
Abstract
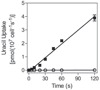
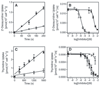
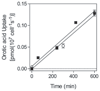
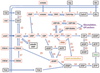
African trypanosomes are capable of both pyrimidine biosynthesis and salvage of preformed pyrimidines from the host. However, uptake of pyrimidines in bloodstream form trypanosomes has not been investigated, making it difficult to judge the relative importance of salvage and synthesis or to design a pyrimidine-based chemotherapy. Detailed characterization of pyrimidine transport activities in bloodstream form Trypanosoma brucei brucei found that these cells express a high-affinity uracil transporter (designated TbU3) that is clearly distinct from the procyclic pyrimidine transporters. This transporter had low affinity for uridine and 2'deoxyuridine and was the sole pyrimidine transporter expressed in these cells. In addition, thymidine was taken up inefficiently through a P1-type nucleoside transporter. Of importance, the anticancer drug 5-fluorouracil was an excellent substrate for TbU3, and several 5-fluoropyrimidine analogs were investigated for uptake and trypanocidal activity; 5F-orotic acid, 5F-2'deoxyuridine displayed activity in the low micromolar range. The metabolism and mode of action of these analogs was determined using metabolomic assessments of T. brucei clonal lines adapted to high levels of these pyrimidine analogs, and of the sensitive parental strains. The analysis showed that 5-fluorouracil is incorporated into a large number of metabolites but likely exerts toxicity through incorporation into RNA. 5F-2'dUrd and 5F-2'dCtd are not incorporated into nucleic acids but act as prodrugs by inhibiting thymidylate synthase as 5F-dUMP. We present the most complete model of pyrimidine salvage in T. brucei to date, supported by genome-wide profiling of the predicted pyrimidine biosynthesis and conversion enzymes.
Figures






References
-
- Al-Salabi MI, Wallace LJM, Lüscher A, Mäser P, Candlish D, Rodenko B, Gould MK, Jabeen I, Ajith SN, De Koning HP. Molecular interactions underlying the unusually high adenosine affinity of a novel Trypanosoma brucei nucleoside transporter. Mol Pharmacol. 2007;71:921–929. - PubMed
-
- Baird JK. Effectiveness of antimalarial drugs. N Engl J Med. 2005;352:1565–1577. - PubMed
-
- Bellofatto V. Pyrimidine transport activities in trypanosomes. Trends Parasitol. 2007;23:187–189. discussion 190. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

