KEGG OC: a large-scale automatic construction of taxonomy-based ortholog clusters
- PMID: 23193276
- PMCID: PMC3531156
- DOI: 10.1093/nar/gks1239
KEGG OC: a large-scale automatic construction of taxonomy-based ortholog clusters
Abstract
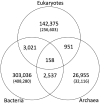
The identification of orthologous genes in an increasing number of fully sequenced genomes is a challenging issue in recent genome science. Here we present KEGG OC (http://www.genome.jp/tools/oc/), a novel database of ortholog clusters (OCs). The current version of KEGG OC contains 1 176 030 OCs, obtained by clustering 8 357 175 genes in 2112 complete genomes (153 eukaryotes, 1830 bacteria and 129 archaea). The OCs were constructed by applying the quasi-clique-based clustering method to all possible protein coding genes in all complete genomes, based on their amino acid sequence similarities. It is computationally efficient to calculate OCs, which enables to regularly update the contents. KEGG OC has the following two features: (i) It consists of all complete genomes of a wide variety of organisms from three domains of life, and the number of organisms is the largest among the existing databases; and (ii) It is compatible with the KEGG database by sharing the same sets of genes and identifiers, which leads to seamless integration of OCs with useful components in KEGG such as biological pathways, pathway modules, functional hierarchy, diseases and drugs. The KEGG OC resources are accessible via OC Viewer that provides an interactive visualization of OCs at different taxonomic levels.
Figures

Similar articles
-
Clusters of orthologous genes for 41 archaeal genomes and implications for evolutionary genomics of archaea.Biol Direct. 2007 Nov 27;2:33. doi: 10.1186/1745-6150-2-33. Biol Direct. 2007. PMID: 18042280 Free PMC article.
-
Automatic detection of conserved gene clusters in multiple genomes by graph comparison and P-quasi grouping.Nucleic Acids Res. 2000 Oct 15;28(20):4029-36. doi: 10.1093/nar/28.20.4029. Nucleic Acids Res. 2000. PMID: 11024184 Free PMC article.
-
KEGG for taxonomy-based analysis of pathways and genomes.Nucleic Acids Res. 2023 Jan 6;51(D1):D587-D592. doi: 10.1093/nar/gkac963. Nucleic Acids Res. 2023. PMID: 36300620 Free PMC article.
-
Comparative Genomics for Prokaryotes.Methods Mol Biol. 2018;1704:55-78. doi: 10.1007/978-1-4939-7463-4_3. Methods Mol Biol. 2018. PMID: 29277863 Review.
-
Genomics of bacteria and archaea: the emerging dynamic view of the prokaryotic world.Nucleic Acids Res. 2008 Dec;36(21):6688-719. doi: 10.1093/nar/gkn668. Epub 2008 Oct 23. Nucleic Acids Res. 2008. PMID: 18948295 Free PMC article. Review.
Cited by
-
Differentially expressed genes between systemic sclerosis and rheumatoid arthritis.Hereditas. 2019 Jun 4;156:17. doi: 10.1186/s41065-019-0091-y. eCollection 2019. Hereditas. 2019. PMID: 31178673 Free PMC article.
-
Functional Properties of Circulating Exosomes Mediated by Surface-Attached Plasma Proteins.J Hematol. 2018 Dec;7(4):149-153. doi: 10.14740/jh412w. Epub 2018 Nov 22. J Hematol. 2018. PMID: 32300430 Free PMC article.
-
Characterising PvRBSA: an exclusive protein from Plasmodium species infecting reticulocytes.Parasit Vectors. 2017 May 18;10(1):243. doi: 10.1186/s13071-017-2185-6. Parasit Vectors. 2017. PMID: 28521840 Free PMC article.
-
Data aggregation at the level of molecular pathways improves stability of experimental transcriptomic and proteomic data.Cell Cycle. 2017 Oct 2;16(19):1810-1823. doi: 10.1080/15384101.2017.1361068. Epub 2017 Aug 21. Cell Cycle. 2017. PMID: 28825872 Free PMC article.
-
Differentially expressed proteins in glioblastoma multiforme identified with a nanobody-based anti-proteome approach and confirmed by OncoFinder as possible tumor-class predictive biomarker candidates.Oncotarget. 2017 Jul 4;8(27):44141-44158. doi: 10.18632/oncotarget.17390. Oncotarget. 2017. PMID: 28498803 Free PMC article.
References
-
- Tatusov RL, Koonin EV, Lipman DJ. A genomic perspective on protein families. Science. 1997;278:631–637. - PubMed

