Folding of large multidomain proteins by partial encapsulation in the chaperonin TRiC/CCT
- PMID: 23197838
- PMCID: PMC3535605
- DOI: 10.1073/pnas.1218836109
Folding of large multidomain proteins by partial encapsulation in the chaperonin TRiC/CCT
Abstract
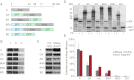
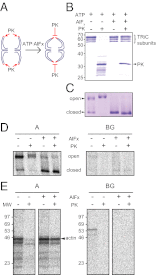
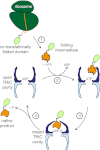
The eukaryotic chaperonin, TRiC/CCT (TRiC, TCP-1 ring complex; CCT, chaperonin containing TCP-1), uses a built-in lid to mediate protein folding in an enclosed central cavity. Recent structural data suggest an effective size limit for the TRiC folding chamber of ∼70 kDa, but numerous chaperonin substrates are substantially larger. Using artificial fusion constructs with actin, an obligate chaperonin substrate, we show that TRiC can mediate folding of large proteins by segmental or domain-wise encapsulation. Single or multiple protein domains up to ∼70 kDa are stably enclosed by stabilizing the ATP-hydrolysis transition state of TRiC. Additional domains, connected by flexible linkers that pass through the central opening of the folding chamber, are excluded and remain accessible to externally added protease. Experiments with the physiological TRiC substrate hSnu114, a 109-kDa multidomain protein, suggest that TRiC has the ability to recognize domain boundaries in partially folded intermediates. In the case of hSnu114, this allows the selective encapsulation of the C-terminal ∼45-kDa domain and segments thereof, presumably reflecting a stepwise folding mechanism. The capacity of the eukaryotic chaperonin to overcome the size limitation of the folding chamber may have facilitated the explosive expansion of the multidomain proteome in eukaryotes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
The structural basis of eukaryotic chaperonin TRiC/CCT: Action and folding.Mol Cells. 2024 Mar;47(3):100012. doi: 10.1016/j.mocell.2024.100012. Epub 2024 Jan 26. Mol Cells. 2024. PMID: 38280673 Free PMC article. Review.
-
TRiC/CCT Chaperonin: Structure and Function.Subcell Biochem. 2019;93:625-654. doi: 10.1007/978-3-030-28151-9_19. Subcell Biochem. 2019. PMID: 31939165 Review.
-
Structural visualization of the tubulin folding pathway directed by human chaperonin TRiC/CCT.Cell. 2022 Dec 8;185(25):4770-4787.e20. doi: 10.1016/j.cell.2022.11.014. Cell. 2022. PMID: 36493755 Free PMC article.
-
Snapshots of actin and tubulin folding inside the TRiC chaperonin.Nat Struct Mol Biol. 2022 May;29(5):420-429. doi: 10.1038/s41594-022-00755-1. Epub 2022 Apr 21. Nat Struct Mol Biol. 2022. PMID: 35449234 Free PMC article.
-
The Mechanism and Function of Group II Chaperonins.J Mol Biol. 2015 Sep 11;427(18):2919-30. doi: 10.1016/j.jmb.2015.04.013. Epub 2015 Apr 30. J Mol Biol. 2015. PMID: 25936650 Free PMC article. Review.
Cited by
-
A Short Tale of the Origin of Proteins and Ribosome Evolution.Microorganisms. 2022 Oct 26;10(11):2115. doi: 10.3390/microorganisms10112115. Microorganisms. 2022. PMID: 36363706 Free PMC article. Review.
-
Novel Binding Partners for CCT and PhLP1 Suggest a Common Folding Mechanism for WD40 Proteins with a 7-Bladed Beta-Propeller Structure.Proteomes. 2021 Oct 2;9(4):40. doi: 10.3390/proteomes9040040. Proteomes. 2021. PMID: 34698247 Free PMC article.
-
Genetic analysis of DEFECTIVE KERNEL1 loop function in three-dimensional body patterning in Physcomitrella patens.Plant Physiol. 2014 Oct;166(2):903-19. doi: 10.1104/pp.114.243758. Epub 2014 Sep 2. Plant Physiol. 2014. PMID: 25185121 Free PMC article.
-
Heat shock protein 104 (HSP104) chaperones soluble Tau via a mechanism distinct from its disaggregase activity.J Biol Chem. 2019 Mar 29;294(13):4956-4965. doi: 10.1074/jbc.RA118.005980. Epub 2019 Feb 4. J Biol Chem. 2019. PMID: 30718279 Free PMC article.
-
Expression and diagnostic value of CCT3 and IQGAP3 in hepatocellular carcinoma.Cancer Cell Int. 2016 Jul 7;16:55. doi: 10.1186/s12935-016-0332-3. eCollection 2016. Cancer Cell Int. 2016. PMID: 27390551 Free PMC article.
References
-
- Ellis RJ. Protein folding: Importance of the Anfinsen cage. Curr Biol. 2003;13(22):R881–R883. - PubMed
-
- Hartl FU. Molecular chaperones in cellular protein folding. Nature. 1996;381(6583):571–579. - PubMed
-
- Horwich AL, Fenton WA, Chapman E, Farr GW. Two families of chaperonin: Physiology and mechanism. Annu Rev Cell Dev Biol. 2007;23:115–145. - PubMed
-
- Frydman J. Folding of newly translated proteins in vivo: The role of molecular chaperones. Annu Rev Biochem. 2001;70:603–647. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

