Chromatin decondensation and nuclear softening accompany Nanog downregulation in embryonic stem cells
- PMID: 23200040
- PMCID: PMC3512036
- DOI: 10.1016/j.bpj.2012.10.015
Chromatin decondensation and nuclear softening accompany Nanog downregulation in embryonic stem cells
Abstract
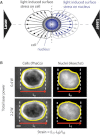
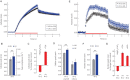
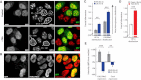
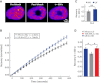
The interplay between epigenetic modification and chromatin compaction is implicated in the regulation of gene expression, and it comprises one of the most fascinating frontiers in cell biology. Although a complete picture is still lacking, it is generally accepted that the differentiation of embryonic stem (ES) cells is accompanied by a selective condensation into heterochromatin with concomitant gene silencing, leaving access only to lineage-specific genes in the euchromatin. ES cells have been reported to have less condensed chromatin, as they are capable of differentiating into any cell type. However, pluripotency itself-even prior to differentiation-is a split state comprising a naïve state and a state in which ES cells prime for differentiation. Here, we show that naïve ES cells decondense their chromatin in the course of downregulating the pluripotency marker Nanog before they initiate lineage commitment. We used fluorescence recovery after photobleaching, and histone modification analysis paired with a novel, to our knowledge, optical stretching method, to show that ES cells in the naïve state have a significantly stiffer nucleus that is coupled to a globally more condensed chromatin state. We link this biophysical phenotype to coinciding epigenetic differences, including histone methylation, and show a strong correlation of chromatin condensation and nuclear stiffness with the expression of Nanog. Besides having implications for transcriptional regulation and embryonic cell sorting and suggesting a putative mechanosensing mechanism, the physical differences point to a system-level regulatory role of chromatin in maintaining pluripotency in embryonic development.
Copyright © 2012 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
References
-
- Chambers I., Colby D., Smith A. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell. 2003;113:643–655. - PubMed
-
- Chambers I., Silva J., Smith A. Nanog safeguards pluripotency and mediates germline development. Nature. 2007;450:1230–1234. - PubMed
-
- Nichols J., Smith A. Naive and primed pluripotent states. Cell Stem Cell. 2009;4:487–492. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

