Xenobiotic-sensing nuclear receptors involved in drug metabolism: a structural perspective
- PMID: 23210723
- PMCID: PMC4878694
- DOI: 10.3109/03602532.2012.740049
Xenobiotic-sensing nuclear receptors involved in drug metabolism: a structural perspective
Abstract
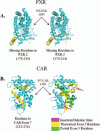
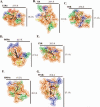
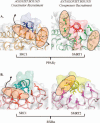
Xenobiotic compounds undergo a critical range of biotransformations performed by the phase I, II, and III drug-metabolizing enzymes. The oxidation, conjugation, and transportation of potentially harmful xenobiotic and endobiotic compounds achieved by these catalytic systems are significantly regulated, at the gene expression level, by members of the nuclear receptor (NR) family of ligand-modulated transcription factors. Activation of NRs by a variety of endo- and exogenous chemicals are elemental to induction and repression of drug-metabolism pathways. The master xenobiotic sensing NRs, the promiscuous pregnane X receptor and less-promiscuous constitutive androstane receptor are crucial to initial ligand recognition, jump-starting the metabolic process. Other receptors, including farnesoid X receptor, vitamin D receptor, hepatocyte nuclear factor 4 alpha, peroxisome proliferator activated receptor, glucocorticoid receptor, liver X receptor, and RAR-related orphan receptor, are not directly linked to promiscuous xenobiotic binding, but clearly play important roles in the modulation of metabolic gene expression. Crystallographic studies of the ligand-binding domains of nine NRs involved in drug metabolism provide key insights into ligand-based and constitutive activity, coregulator recruitment, and gene regulation. Structures of other, noncanonical transcription factors also shed light on secondary, but important, pathways of control. Pharmacological targeting of some of these nuclear and atypical receptors has been instituted as a means to treat metabolic and developmental disorders and provides a future avenue to be explored for other members of the xenobiotic-sensing NRs.
Figures





Similar articles
-
Induction of phase I, II and III drug metabolism/transport by xenobiotics.Arch Pharm Res. 2005 Mar;28(3):249-68. doi: 10.1007/BF02977789. Arch Pharm Res. 2005. PMID: 15832810 Review.
-
Regulation of drug-metabolizing enzymes by xenobiotic receptors: PXR and CAR.Adv Drug Deliv Rev. 2010 Oct 30;62(13):1238-49. doi: 10.1016/j.addr.2010.08.006. Epub 2010 Aug 17. Adv Drug Deliv Rev. 2010. PMID: 20727377 Free PMC article. Review.
-
Xenobiotic receptor humanized mice and their utility.Drug Metab Rev. 2013 Feb;45(1):110-21. doi: 10.3109/03602532.2012.738687. Epub 2012 Nov 23. Drug Metab Rev. 2013. PMID: 23173549 Review.
-
Nuclear receptors in bile acid metabolism.Drug Metab Rev. 2013 Feb;45(1):145-55. doi: 10.3109/03602532.2012.740048. Drug Metab Rev. 2013. PMID: 23330546 Free PMC article. Review.
-
Beyond CAR and PXR.Curr Drug Metab. 2005 Aug;6(4):385-97. doi: 10.2174/1389200054633907. Curr Drug Metab. 2005. PMID: 16101576 Review.
Cited by
-
Nuclear hormone receptors promote gut and glia detoxifying enzyme induction and protect C. elegans from the mold P. brevicompactum.Cell Rep. 2021 Dec 28;37(13):110166. doi: 10.1016/j.celrep.2021.110166. Cell Rep. 2021. PMID: 34965433 Free PMC article.
-
Xenobiotic metabolism and transport in Caenorhabditis elegans.J Toxicol Environ Health B Crit Rev. 2021 Feb 17;24(2):51-94. doi: 10.1080/10937404.2021.1884921. Epub 2021 Feb 22. J Toxicol Environ Health B Crit Rev. 2021. PMID: 33616007 Free PMC article. Review.
-
Bile Acids Promote Hepatic Biotransformation and Excretion of Aflatoxin B1 in Broiler Chickens.Toxins (Basel). 2023 Dec 9;15(12):694. doi: 10.3390/toxins15120694. Toxins (Basel). 2023. PMID: 38133198 Free PMC article.
-
Changes in the Gut Microbiota May Affect the Clinical Efficacy of Oral Anticoagulants.Front Pharmacol. 2022 Mar 25;13:860237. doi: 10.3389/fphar.2022.860237. eCollection 2022. Front Pharmacol. 2022. PMID: 35401180 Free PMC article.
-
Nuclear Receptor Metabolism of Bile Acids and Xenobiotics: A Coordinated Detoxification System with Impact on Health and Diseases.Int J Mol Sci. 2018 Nov 17;19(11):3630. doi: 10.3390/ijms19113630. Int J Mol Sci. 2018. PMID: 30453651 Free PMC article. Review.
References
-
- Aleksunes LM, Manautou JE. Emerging role of Nrf2 in protecting against hepatic and gastrointestinal disease. Toxicol Pathol. 2007;35:459–473. - PubMed
-
- Alnouti Y, Klaassen CD. Regulation of sulfotransferase enzymes by prototypical microsomal enzyme inducers in mice. J Pharmacol Exp Ther. 2008;324:612–621. - PubMed
-
- André E, Gawlas K, Becker-André M. A novel isoform of the orphan nuclear receptor RORbeta is specifically expressed in pineal gland and retina. Gene. 1998;216:277–283. - PubMed
-
- Arita K, Nanda A, Wessagowit V, Akiyama M, Alsaleh QA, McGrath JA. A novel mutation in the VDR gene in hereditary vitamin D-resistant rickets. Br J Dermatol. 2008;158:168–171. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
