Mechanism of CREB recognition and coactivation by the CREB-regulated transcriptional coactivator CRTC2
- PMID: 23213254
- PMCID: PMC3529076
- DOI: 10.1073/pnas.1219028109
Mechanism of CREB recognition and coactivation by the CREB-regulated transcriptional coactivator CRTC2
Abstract
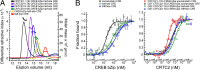
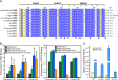
Basic leucine zipper (bZip) transcription factors regulate cellular gene expression in response to a variety of extracellular signals and nutrient cues. Although the bZip domain is widely known to play significant roles in DNA binding and dimerization, recent studies point to an additional role for this motif in the recruitment of the transcriptional apparatus. For example, the cAMP response element binding protein (CREB)-regulated transcriptional coactivator (CRTC) family of transcriptional coactivators has been proposed to promote the expression of calcium and cAMP responsive genes, by binding to the CREB bZip in response to extracellular signals. Here we show that the CREB-binding domain (CBD) of CRTC2 folds into a single isolated 28-residue helix that seems to be critical for its interaction with the CREB bZip. The interaction is of micromolar affinity on palindromic and variant half-site cAMP response elements (CREs). The CBD and CREB assemble on the CRE with 2:2:1 stoichiometry, consistent with the presence of one CRTC binding site on each CREB monomer. Indeed, the CBD helix and the solvent-exposed residues in the dimeric CREB bZip coiled-coil form an extended protein-protein interface. Because mutation of relevant bZip residues in this interface disrupts the CRTC interaction without affecting DNA binding, our results illustrate that distinct DNA binding and transactivation functions are encoded within the structural constraints of a canonical bZip domain.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


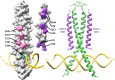
References
-
- Vinson C, Acharya A, Taparowsky EJ. Deciphering B-ZIP transcription factor interactions in vitro and in vivo. Biochim Biophys Acta. 2006;1759(1-2):4–12. - PubMed
-
- Montminy M. Transcriptional regulation by cyclic AMP. Annu Rev Biochem. 1997;66:807–822. - PubMed
-
- Chrivia JC, et al. Phosphorylated CREB binds specifically to the nuclear protein CBP. Nature. 1993;365(6449):855–859. - PubMed
-
- Conkright MD, et al. TORCs: Transducers of regulated CREB activity. Mol Cell. 2003;12(2):413–423. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

