Taperin (c9orf75), a mutated gene in nonsyndromic deafness, encodes a vertebrate specific, nuclear localized protein phosphatase one alpha (PP1α) docking protein
- PMID: 23213405
- PMCID: PMC3507197
- DOI: 10.1242/bio.2011049
Taperin (c9orf75), a mutated gene in nonsyndromic deafness, encodes a vertebrate specific, nuclear localized protein phosphatase one alpha (PP1α) docking protein
Abstract
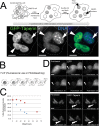
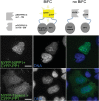
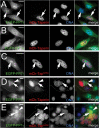
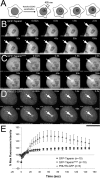
The promiscuous activity of protein phosphatase one (PP1) is controlled in the cell by associated proteins termed regulatory or targeting subunits. Using biochemical and proteomic approaches we demonstrate that the autosomal recessive nonsyndromic hearing loss gene, taperin (C9orf75), encodes a protein that preferentially docks the alpha isoform of PP1. Taperin associates with PP1 through a classic 'RVxF' motif and suppresses the general phosphatase activity of the enzyme. The steady-state localization of taperin is predominantly nuclear, however we demonstrate here that the protein can shuttle between the nucleus and cytoplasm and that it is found complexed to PP1 in both of these cellular compartments. Although originally identified as a hearing loss gene, Western blot analyses with taperin-specific antibodies revealed that the protein is widely expressed across mammalian tissues as multiple splice variants. Taperin is a recent proteome addition appearing during the vertebrate lineage with the PP1 binding site embedded within the most conserved region of the protein. Taperin also shares an ancestral relationship with the cytosolic actin binding protein phostensin, another PP1 interacting partner. Quantitative Stable Isotope Labeling by Amino acids in Culture (SILAC)-based mass spectrometry was employed to uncover additional taperin binding partners, and revealed an interaction with the DNA damage response proteins Ku70, Ku80, PARP and topoisomerases I and IIα. Consistent with this, we demonstrate the active recruitment of taperin to sites of DNA damage. This makes taperin a new addition to the family of PP1 targeting subunits involved in the DNA damage repair pathway.
Keywords: DNA damage; Nonsyndromic deafness; Nucleus; Phostensin; Protein phosphatase one (PP1); Protein phosphorylation.
Figures



References
-
- Bespalova I. N., Van Camp G., Bom S. J., Brown D. J., Cryns K., DeWan A. T., Erson A. E., Flothmann K., Kunst H. P., Kurnool P. et al. (2001). Mutations in the Wolfram syndrome 1 gene (WFS1) are a common cause of low frequency sensorineural hearing loss. Hum. Mol. Genet. 10, 2501–2508 10.1093/hmg/10.22.2501 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

