The state of the human proteome in 2012 as viewed through PeptideAtlas
- PMID: 23215161
- PMCID: PMC3928036
- DOI: 10.1021/pr301012j
The state of the human proteome in 2012 as viewed through PeptideAtlas
Abstract
The Human Proteome Project was launched in September 2010 with the goal of characterizing at least one protein product from each protein-coding gene. Here we assess how much of the proteome has been detected to date via tandem mass spectrometry by analyzing PeptideAtlas, a compendium of human derived LC-MS/MS proteomics data from many laboratories around the world. All data sets are processed with a consistent set of parameters using the Trans-Proteomic Pipeline and subjected to a 1% protein FDR filter before inclusion in PeptideAtlas. Therefore, PeptideAtlas contains only high confidence protein identifications. To increase proteome coverage, we explored new comprehensive public data sources for data likely to add new proteins to the Human PeptideAtlas. We then folded these data into a Human PeptideAtlas 2012 build and mapped it to Swiss-Prot, a protein sequence database curated to contain one entry per human protein coding gene. We find that this latest PeptideAtlas build includes at least one peptide for each of ~12500 Swiss-Prot entries, leaving ~7500 gene products yet to be confidently cataloged. We characterize these "PA-unseen" proteins in terms of tissue localization, transcript abundance, and Gene Ontology enrichment, and propose reasons for their absence from PeptideAtlas and strategies for detecting them in the future.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
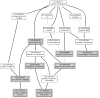

References
-
- Paik YK, Jeong SK, Omenn GS, Uhlen M, Hanash S, Cho SY, Lee HJ, Na K, Choi EY, Yan F, Zhang F, Zhang Y, Snyder M, Cheng Y, Chen R, Marko-Varga G, Deutsch EW, Kim H, Kwon JY, Aebersold R, Bairoch A, Taylor AD, Kim KY, Lee EY, Hochstrasser D, Legrain P, Hancock WS. The Chromosome-Centric Human Proteome Project for cataloging proteins encoded in the genome. Nat Biotechnol. 2012;30(3):221–3. - PubMed
-
- Legrain P, Aebersold R, Archakov A, Bairoch A, Bala K, Beretta L, Bergeron J, Borchers CH, Corthals GL, Costello CE, Deutsch EW, Domon B, Hancock W, He F, Hochstrasser D, Marko-Varga G, Salekdeh GH, Sechi S, Snyder M, Srivastava S, Uhlen M, Wu CH, Yamamoto T, Paik YK, Omenn GS. The human proteome project: current state and future direction. Mol Cell Proteomics. 2011;10(7):M111 009993. - PMC - PubMed
-
- Flicek P, Amode MR, Barrell D, Beal K, Brent S, Carvalho-Silva D, Clapham P, Coates G, Fairley S, Fitzgerald S, Gil L, Gordon L, Hendrix M, Hourlier T, Johnson N, Kahari AK, Keefe D, Keenan S, Kinsella R, Komorowska M, Koscielny G, Kulesha E, Larsson P, Longden I, McLaren W, Muffato M, Overduin B, Pignatelli M, Pritchard B, Riat HS, Ritchie GR, Ruffier M, Schuster M, Sobral D, Tang YA, Taylor K, Trevanion S, Vandrovcova J, White S, Wilson M, Wilder SP, Aken BL, Birney E, Cunningham F, Dunham I, Durbin R, Fernandez-Suarez XM, Harrow J, Herrero J, Hubbard TJ, Parker A, Proctor G, Spudich G, Vogel J, Yates A, Zadissa A, Searle SM. Ensembl 2012. Nucleic Acids Res. 2012;40(Database issue):D84–90. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

