A heme-sensing mechanism in the translational regulation of mitochondrial cytochrome c oxidase biogenesis
- PMID: 23217259
- PMCID: PMC3523284
- DOI: 10.1016/j.cmet.2012.10.018
A heme-sensing mechanism in the translational regulation of mitochondrial cytochrome c oxidase biogenesis
Abstract
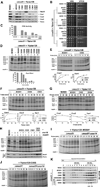
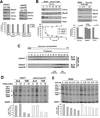
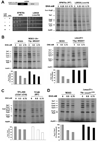
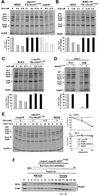
Heme plays fundamental roles as cofactor and signaling molecule in multiple pathways devoted to oxygen sensing and utilization in aerobic organisms. For cellular respiration, heme serves as a prosthetic group in electron transfer proteins and redox enzymes. Here we report that in the yeast Saccharomyces cerevisiae, a heme-sensing mechanism translationally controls the biogenesis of cytochrome c oxidase (COX), the terminal mitochondrial respiratory chain enzyme. We show that Mss51, a COX1 mRNA-specific translational activator and Cox1 chaperone, which coordinates Cox1 synthesis in mitoribosomes with its assembly in COX, is a heme-binding protein. Mss51 contains two heme regulatory motifs or Cys-Pro-X domains located in its N terminus. Using a combination of in vitro and in vivo approaches, we have demonstrated that these motifs are important for heme binding and efficient performance of Mss51 functions. We conclude that heme sensing by Mss51 regulates COX biogenesis and aerobic energy production.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures


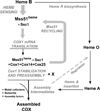
References
-
- Barros MH, Tzagoloff A. Regulation of the heme A biosynthetic pathway in Saccharomyces cerevisiae. FEBS Lett. 2002;516:119–123. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

