Affinofile profiling: how efficiency of CD4/CCR5 usage impacts the biological and pathogenic phenotype of HIV
- PMID: 23217618
- PMCID: PMC3522187
- DOI: 10.1016/j.virol.2012.09.043
Affinofile profiling: how efficiency of CD4/CCR5 usage impacts the biological and pathogenic phenotype of HIV
Abstract
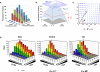
HIV-1 envelope (Env) uses CD4 and a coreceptor (CCR5 and/or CXCR4) for viral entry. The efficiency of receptor/coreceptor mediated entry has important implications for HIV pathogenesis and transmission. The advent of CCR5 inhibitors in clinical use also underscores the need for quantitative and predictive tools that can guide therapeutic management. Historically, measuring the efficiency of CD4/CCR5 mediated HIV entry has relied on surrogate and relatively slow throughput assays that cannot adequately capture the full spectrum of Env phenotypes. In this review, we discuss the details of the Affinofile receptor affinity profiling system that has provided a quantitative and higher throughput method to characterize viral entry efficiency as a function of CD4 and CCR5 expression levels. We will then review how the Affinofile system has been used to reveal the distinct pathophysiological properties associated with Env entry phenotypes and discuss potential shortcomings of the current system.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Distinct HIV-1 entry phenotypes are associated with transmission, subtype specificity, and resistance to broadly neutralizing antibodies.Retrovirology. 2014 Jun 23;11:48. doi: 10.1186/1742-4690-11-48. Retrovirology. 2014. PMID: 24957778 Free PMC article.
-
Quantifying CD4/CCR5 Usage Efficiency of HIV-1 Env Using the Affinofile System.Methods Mol Biol. 2016;1354:3-20. doi: 10.1007/978-1-4939-3046-3_1. Methods Mol Biol. 2016. PMID: 26714701
-
A double-mimetic peptide efficiently neutralizes HIV-1 by bridging the CD4- and coreceptor-binding sites of gp120.J Virol. 2014 Mar;88(6):3353-8. doi: 10.1128/JVI.03800-13. Epub 2014 Jan 3. J Virol. 2014. PMID: 24390333 Free PMC article.
-
Effect of HIV-1 subtype and tropism on treatment with chemokine coreceptor entry inhibitors; overview of viral entry inhibition.Crit Rev Microbiol. 2015;41(4):473-87. doi: 10.3109/1040841X.2013.867829. Epub 2014 Mar 17. Crit Rev Microbiol. 2015. PMID: 24635642 Review.
-
How HIV changes its tropism: evolution and adaptation?Curr Opin HIV AIDS. 2009 Mar;4(2):125-30. doi: 10.1097/COH.0b013e3283223d61. Curr Opin HIV AIDS. 2009. PMID: 19339951 Free PMC article. Review.
Cited by
-
Sendai virus, an RNA virus with no risk of genomic integration, delivers CRISPR/Cas9 for efficient gene editing.Mol Ther Methods Clin Dev. 2016 Aug 24;3:16057. doi: 10.1038/mtm.2016.57. eCollection 2016. Mol Ther Methods Clin Dev. 2016. PMID: 27606350 Free PMC article.
-
A single-residue change in the HIV-1 V3 loop associated with maraviroc resistance impairs CCR5 binding affinity while increasing replicative capacity.Retrovirology. 2015 Jun 18;12:50. doi: 10.1186/s12977-015-0177-1. Retrovirology. 2015. PMID: 26081316 Free PMC article.
-
Modular Lentiviral Vectors for Highly Efficient Transgene Expression in Resting Immune Cells.Viruses. 2021 Jun 18;13(6):1170. doi: 10.3390/v13061170. Viruses. 2021. PMID: 34207354 Free PMC article.
-
Regulation of epitope exposure in the gp41 membrane-proximal external region through interactions at the apex of HIV-1 Env.PLoS Pathog. 2022 May 18;18(5):e1010531. doi: 10.1371/journal.ppat.1010531. eCollection 2022 May. PLoS Pathog. 2022. PMID: 35584191 Free PMC article.
-
The Effects of Statistical Multiplicity of Infection on Virus Quantification and Infectivity Assays.Biophys J. 2018 Jun 19;114(12):2974-2985. doi: 10.1016/j.bpj.2018.05.005. Biophys J. 2018. PMID: 29925033 Free PMC article.
References
-
- Berger EA, Doms RW, Fenyo EM, Korber BT, Littman DR, Moore JP, Sattentau QJ, Schuitemaker H, Sodroski J, Weiss RA. A new classification for HIV-1. Nature. 1998;391:240. - PubMed
-
- Blaak H, Ran LJ, Rientsma R, Schuitemaker H. Susceptibility of in vitro stimulated PBMC to infection with NSI HIV-1 is associated with levels of CCR5 expression and beta-chemokine production. Virology. 2000;267:237–46. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

