The chloroplast genome of Pellia endiviifolia: gene content, RNA-editing pattern, and the origin of chloroplast editing
- PMID: 23221608
- PMCID: PMC3542565
- DOI: 10.1093/gbe/evs114
The chloroplast genome of Pellia endiviifolia: gene content, RNA-editing pattern, and the origin of chloroplast editing
Abstract
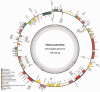
RNA editing is a post-transcriptional process that can act upon transcripts from mitochondrial, nuclear, and chloroplast genomes. In chloroplasts, single-nucleotide conversions in mRNAs via RNA editing occur at different frequencies across the plant kingdom. These range from several hundred edited sites in some mosses and ferns to lower frequencies in seed plants and the complete lack of RNA editing in the liverwort Marchantia polymorpha. Here, we report the sequence and edited sites of the chloroplast genome from the liverwort Pellia endiviifolia. The type and frequency of chloroplast RNA editing display a pattern highly similar to that in seed plants. Analyses of the C to U conversions and the genomic context in which the editing sites are embedded provide evidence in favor of the hypothesis that chloroplast RNA editing evolved to compensate mutations in the first land plants.
Figures
Similar articles
-
The liverwort Pellia endiviifolia shares microtranscriptomic traits that are common to green algae and land plants.New Phytol. 2015 Apr;206(1):352-367. doi: 10.1111/nph.13220. Epub 2014 Dec 19. New Phytol. 2015. PMID: 25530158 Free PMC article.
-
The identification of differentially expressed genes in male and female gametophytes of simple thalloid liverwort Pellia endiviifolia sp. B using an RNA-seq approach.Planta. 2020 Jul 15;252(2):21. doi: 10.1007/s00425-020-03424-z. Planta. 2020. PMID: 32671488 Free PMC article.
-
Reverse U-to-C editing exceeds C-to-U RNA editing in some ferns - a monilophyte-wide comparison of chloroplast and mitochondrial RNA editing suggests independent evolution of the two processes in both organelles.BMC Evol Biol. 2016 Jun 21;16(1):134. doi: 10.1186/s12862-016-0707-z. BMC Evol Biol. 2016. PMID: 27329857 Free PMC article.
-
Chloroplast and mitochondrial genomes from a liverwort, Marchantia polymorpha--gene organization and molecular evolution.Biosci Biotechnol Biochem. 1996 Jan;60(1):16-24. doi: 10.1271/bbb.60.16. Biosci Biotechnol Biochem. 1996. PMID: 8824820 Review.
-
Gene content, organization and molecular evolution of plant organellar genomes and sex chromosomes: insights from the case of the liverwort Marchantia polymorpha.Proc Jpn Acad Ser B Phys Biol Sci. 2009;85(3):108-24. doi: 10.2183/pjab.85.108. Proc Jpn Acad Ser B Phys Biol Sci. 2009. PMID: 19282647 Free PMC article. Review.
Cited by
-
The complete plastid genome sequence of the enigmatic moss, Takakia lepidozioides (Takakiopsida, Bryophyta): evolutionary perspectives on the largest collection of genes in mosses and the intensive RNA editing.Plant Mol Biol. 2021 Nov;107(4-5):431-449. doi: 10.1007/s11103-021-01214-z. Epub 2021 Nov 24. Plant Mol Biol. 2021. PMID: 34817767
-
Complete Chloroplast Genome Sequence and Phylogenetic Analysis of Aster tataricus.Molecules. 2018 Sep 21;23(10):2426. doi: 10.3390/molecules23102426. Molecules. 2018. PMID: 30248930 Free PMC article.
-
The Increase of Simple Sequence Repeats during Diversification of Marchantiidae, An Early Land Plant Lineage, Leads to the First Known Expansion of Inverted Repeats in the Evolutionarily-Stable Structure of Liverwort Plastomes.Genes (Basel). 2020 Mar 12;11(3):299. doi: 10.3390/genes11030299. Genes (Basel). 2020. PMID: 32178248 Free PMC article.
-
Plant organelle RNA editing and its specificity factors: enhancements of analyses and new database features in PREPACT 3.0.BMC Bioinformatics. 2018 Jul 3;19(1):255. doi: 10.1186/s12859-018-2244-9. BMC Bioinformatics. 2018. PMID: 29970001 Free PMC article.
-
Lycophyte plastid genomics: extreme variation in GC, gene and intron content and multiple inversions between a direct and inverted orientation of the rRNA repeat.New Phytol. 2019 Apr;222(2):1061-1075. doi: 10.1111/nph.15650. Epub 2019 Jan 24. New Phytol. 2019. PMID: 30556907 Free PMC article.
References
-
- Bolte K, et al. Protein targeting into secondary plastids. J Eukaryot Microbiol. 2009;56:9–15. - PubMed
-
- Covello PS, Gray MW. On the evolution of RNA editing. Trends Genet. 1993;9:265–268. - PubMed
-
- Doyle JJ, Doyle JL. Isolation of plant DNA from fresh tissue. Focus. 1990;12:13–15.
-
- Dang Y, Green BR. Substitutional editing of Heterocapsa triquetra chloroplast transcripts and a folding model for its divergent chloroplast 16S rRNA. Gene. 2009;442:73–80. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

