Orthologous gene clusters and taxon signature genes for viruses of prokaryotes
- PMID: 23222723
- PMCID: PMC3571318
- DOI: 10.1128/JB.01801-12
Orthologous gene clusters and taxon signature genes for viruses of prokaryotes
Abstract
Viruses are the most abundant biological entities on earth and encompass a vast amount of genetic diversity. The recent rapid increase in the number of sequenced viral genomes has created unprecedented opportunities for gaining new insight into the structure and evolution of the virosphere. Here, we present an update of the phage orthologous groups (POGs), a collection of 4,542 clusters of orthologous genes from bacteriophages that now also includes viruses infecting archaea and encompasses more than 1,000 distinct virus genomes. Analysis of this expanded data set shows that the number of POGs keeps growing without saturation and that a substantial majority of the POGs remain specific to viruses, lacking homologues in prokaryotic cells, outside known proviruses. Thus, the great majority of virus genes apparently remains to be discovered. A complementary observation is that numerous viral genomes remain poorly, if at all, covered by POGs. The genome coverage by POGs is expected to increase as more genomes are sequenced. Taxon-specific, single-copy signature genes that are not observed in prokaryotic genomes outside detected proviruses were identified for two-thirds of the 57 taxa (those with genomes available from at least 3 distinct viruses), with half of these present in all members of the respective taxon. These signatures can be used to specifically identify the presence and quantify the abundance of viruses from particular taxa in metagenomic samples and thus gain new insights into the ecology and evolution of viruses in relation to their hosts.
Figures
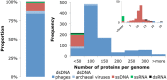
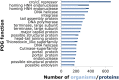
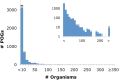

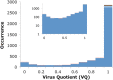
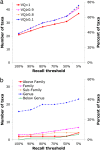
References
-
- Breitbart M, Rohwer F. 2005. Here a virus, there a virus, everywhere the same virus? Trends Microbiol. 13(6):278–284 - PubMed
-
- Suttle CA. 2007. Marine viruses–major players in the global ecosystem. Nat. Rev. Microbiol. 5(10):801–812 - PubMed
-
- Ansorge WJ. 2009. Next-generation DNA sequencing techniques. Nat. Biotechnol. 25(4):195–203 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

