Replication of GWAS-identified neuroblastoma risk loci strengthens the role of BARD1 and affirms the cumulative effect of genetic variations on disease susceptibility
- PMID: 23222812
- PMCID: PMC3716226
- DOI: 10.1093/carcin/bgs380
Replication of GWAS-identified neuroblastoma risk loci strengthens the role of BARD1 and affirms the cumulative effect of genetic variations on disease susceptibility
Erratum in
- Carcinogenesis. 2014 Mar;35(3):737
Abstract
Several neuroblastoma (NB) susceptibility loci have been identified within LINC00340, BARD1, LMO1, DUSP12, HSD17B12, DDX4, IL31RA, HACE1 and LIN28B by genome-wide association (GWA) studies including European American individuals. To validate and comprehensively evaluate the impact of the identified NB variants on disease risk and phenotype, we analyzed 16 single nucleotide polymorphisms (SNPs) in an Italian population (370 cases and 809 controls). We assessed their regulatory activity on gene expression in lymphoblastoid (LCLs) and NB cell lines. We evaluated the cumulative effect of the independent loci on NB risk and high-risk phenotype development in Italian and European American (1627 cases and 2575 controls) populations. All NB susceptibility genes replicated in the Italian dataset except for DDX4 and IL31RA, and the most significant SNP was rs6435862 in BARD1 (P = 8.4 × 10(-15)). BARD1 showed an additional and independent SNP association (rs7585356). This variant influenced BARD1 mRNA expression in LCLs and NB cell lines. No evidence of epistasis among the NB-associated variants was detected, whereas a cumulative effect of risk variants on NB risk (European Americans: P (trend) = 6.9 × 10(-30), Italians: P (trend) = 8.55 × 10(13)) and development of high-risk phenotype (European Americans: P (trend) = 6.9 × 10(-13), Italians: P (trend) = 2.2 × 10(-1)) was observed in a dose-dependent manner. These results provide further evidence that the risk loci identified in GWA studies contribute to NB susceptibility in distinct populations and strengthen the role of BARD1 as major genetic contributor to NB risk. This study shows that even in the absence of interaction the combination of several low-penetrance alleles has potential to distinguish subgroups of patients at different risks of developing NB.
Figures
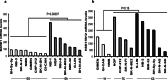

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

