Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine
- PMID: 23223451
- PMCID: PMC3847602
- DOI: 10.1126/science.1229277
Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine
Abstract
Mouse primordial germ cells (PGCs) undergo sequential epigenetic changes and genome-wide DNA demethylation to reset the epigenome for totipotency. Here, we demonstrate that erasure of CpG methylation (5mC) in PGCs occurs via conversion to 5-hydroxymethylcytosine (5hmC), driven by high levels of TET1 and TET2. Global conversion to 5hmC initiates asynchronously among PGCs at embryonic day (E) 9.5 to E10.5 and accounts for the unique process of imprint erasure. Mechanistically, 5hmC enrichment is followed by its protracted decline thereafter at a rate consistent with replication-coupled dilution. The conversion to 5hmC is an important component of parallel redundant systems that drive comprehensive reprogramming in PGCs. Nonetheless, we identify rare regulatory elements that escape systematic DNA demethylation in PGCs, providing a potential mechanistic basis for transgenerational epigenetic inheritance.
Figures

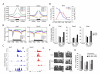


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

