Coral thermal tolerance: tuning gene expression to resist thermal stress
- PMID: 23226355
- PMCID: PMC3511300
- DOI: 10.1371/journal.pone.0050685
Coral thermal tolerance: tuning gene expression to resist thermal stress
Erratum in
- PLoS One. 2013;8(10). doi:10.1371/annotation/2b623dcd-9413-47b0-990d-c46c8e662ad4
Abstract
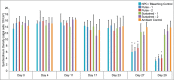
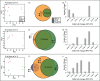
The acclimatization capacity of corals is a critical consideration in the persistence of coral reefs under stresses imposed by global climate change. The stress history of corals plays a role in subsequent response to heat stress, but the transcriptomic changes associated with these plastic changes have not been previously explored. In order to identify host transcriptomic changes associated with acquired thermal tolerance in the scleractinian coral Acropora millepora, corals preconditioned to a sub-lethal temperature of 3°C below bleaching threshold temperature were compared to both non-preconditioned corals and untreated controls using a cDNA microarray platform. After eight days of hyperthermal challenge, conditions under which non-preconditioned corals bleached and preconditioned corals (thermal-tolerant) maintained Symbiodinium density, a clear differentiation in the transcriptional profiles was revealed among the condition examined. Among these changes, nine differentially expressed genes separated preconditioned corals from non-preconditioned corals, with 42 genes differentially expressed between control and preconditioned treatments, and 70 genes between non-preconditioned corals and controls. Differentially expressed genes included components of an apoptotic signaling cascade, which suggest the inhibition of apoptosis in preconditioned corals. Additionally, lectins and genes involved in response to oxidative stress were also detected. One dominant pattern was the apparent tuning of gene expression observed between preconditioned and non-preconditioned treatments; that is, differences in expression magnitude were more apparent than differences in the identity of genes differentially expressed. Our work revealed a transcriptomic signature underlying the tolerance associated with coral thermal history, and suggests that understanding the molecular mechanisms behind physiological acclimatization would be critical for the modeling of reefs in impending climate change scenarios.
Conflict of interest statement
Figures

References
-
- Brander LM, Van Beukering P, Cesar HSJ (2007) The recreational value of coral reefs: a meta-analysis. Ecol Econ 63: 209–218.
-
- Costanza R, dArge R, deGroot R, Farber S, Grasso M, et al. (1997) The value of the world’s ecosystem services and natural capital. Nature 387: 253–260.
-
- Wilkinson C (2004) Status of coral reefs of the World: 2004. Volume 2. Status of coral reefs of the world.
-
- Fitt WK, Brown BE, Warner ME, Dunne RP (2001) Coral bleaching: interpretation of thermal tolerance limits and thermal thresholds in tropical corals. Coral Reefs 20: 51–65.
-
- Glynn PW (1984) Widespread coral mortality and the 1982–83 El Niño warming event. Environ Conserv 11: 133–146.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

