Receptor tyrosine kinase ErbB2 translocates into mitochondria and regulates cellular metabolism
- PMID: 23232401
- PMCID: PMC3521558
- DOI: 10.1038/ncomms2236
Receptor tyrosine kinase ErbB2 translocates into mitochondria and regulates cellular metabolism
Abstract
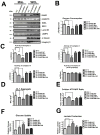
It is well known that ErbB2, a receptor tyrosine kinase, localizes to the plasma membrane. Here we describe a novel observation that ErbB2 also localizes in mitochondria of cancer cells and patient samples. We found that ErbB2 translocates into mitochondria through association with mtHSP70. Additionally, mitochondrial ErbB2 (mtErbB2) negatively regulates mitochondrial respiratory functions. Oxygen consumption and activities of complexes of the mitochondrial electron transport chain were decreased in mtErbB2-overexpressing cells. Mitochondrial membrane potential and cellular ATP levels were also decreased. In contrast, mtErbB2 enhanced cellular glycolysis. The translocation of ErbB2 and its impact on mitochondrial function are kinase dependent. Interestingly, cancer cells with higher levels of mtErbB2 were more resistant to the ErbB2-targeting antibody trastuzumab. Our study provides a novel perspective on the metabolic regulatory function of ErbB2 and reveals that mtErbB2 has an important role in the regulation of cellular metabolism and cancer cell resistance to therapeutics.
Conflict of interest statement
Competing financial interests: The authors declare no competing financial interests.
Figures





Similar articles
-
Hsp90 inhibitor 17-AAG reduces ErbB2 levels and inhibits proliferation of the trastuzumab resistant breast tumor cell line JIMT-1.Immunol Lett. 2006 Apr 15;104(1-2):146-55. doi: 10.1016/j.imlet.2005.11.018. Epub 2005 Dec 12. Immunol Lett. 2006. PMID: 16384610
-
ERBB2 phosphorylation and trastuzumab sensitivity of breast cancer cell lines.Oncogene. 2007 Nov 1;26(50):7163-9. doi: 10.1038/sj.onc.1210528. Epub 2007 May 21. Oncogene. 2007. PMID: 17525746
-
Potent anti-proliferative effects of metformin on trastuzumab-resistant breast cancer cells via inhibition of erbB2/IGF-1 receptor interactions.Cell Cycle. 2011 Sep 1;10(17):2959-66. doi: 10.4161/cc.10.17.16359. Epub 2011 Sep 1. Cell Cycle. 2011. PMID: 21862872
-
Mechanisms of ErbB2-mediated paclitaxel resistance and trastuzumab-mediated paclitaxel sensitization in ErbB2-overexpressing breast cancers.Semin Oncol. 2001 Oct;28(5 Suppl 16):12-7. doi: 10.1016/s0093-7754(01)90277-5. Semin Oncol. 2001. PMID: 11706391 Review.
-
Cardiotoxicity in signal transduction therapeutics: erbB2 antibodies and the heart.Semin Oncol. 2001 Oct;28(5 Suppl 16):18-26. doi: 10.1016/s0093-7754(01)90278-7. Semin Oncol. 2001. PMID: 11706392 Review.
Cited by
-
Melatonin Signaling Pathways Implicated in Metabolic Processes in Human Granulosa Cells (KGN).Int J Mol Sci. 2022 Mar 10;23(6):2988. doi: 10.3390/ijms23062988. Int J Mol Sci. 2022. PMID: 35328408 Free PMC article.
-
Integration of platinum nanoparticles with a volumetric bar-chart chip for biomarker assays.Angew Chem Int Ed Engl. 2014 Nov 10;53(46):12451-5. doi: 10.1002/anie.201404349. Epub 2014 Jul 17. Angew Chem Int Ed Engl. 2014. PMID: 25044863 Free PMC article.
-
Mitochondrial determinants of cancer health disparities.Semin Cancer Biol. 2017 Dec;47:125-146. doi: 10.1016/j.semcancer.2017.05.001. Epub 2017 May 6. Semin Cancer Biol. 2017. PMID: 28487205 Free PMC article. Review.
-
Src drives the Warburg effect and therapy resistance by inactivating pyruvate dehydrogenase through tyrosine-289 phosphorylation.Oncotarget. 2016 May 3;7(18):25113-24. doi: 10.18632/oncotarget.7159. Oncotarget. 2016. PMID: 26848621 Free PMC article.
-
The ERBB2 c.1795C>T, p.Arg599Cys variant is associated with left ventricular outflow tract obstruction defects in humans.HGG Adv. 2025 Jul 10;6(3):100446. doi: 10.1016/j.xhgg.2025.100446. Epub 2025 May 5. HGG Adv. 2025. PMID: 40329538 Free PMC article.
References
-
- Warburg O. On respiratory impairment in cancer cells. Science. 1956;124:269–270. - PubMed
-
- Kim JW, Dang CV. Cancer’s molecular sweet tooth and the Warburg effect. Cancer Res. 2006;66:8927–30. - PubMed
-
- Chen Z, Lu WQ, Garcia-Prieto C, Huang P. The Warburg effect and its cancer therapeutic implications. J Bioenerg Biomembr. 2007;39:267–274. - PubMed
-
- Gatenby RA, Gillies RJ. Glycolysis in cancer: a potential target for therapy. Int J Biochem Cell Biol. 2007;39:1358–66. - PubMed
-
- DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 2008;7:11–20. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

