Chromosome dynamics visualized with an anti-centromeric histone H3 antibody in Allium
- PMID: 23236469
- PMCID: PMC3517398
- DOI: 10.1371/journal.pone.0051315
Chromosome dynamics visualized with an anti-centromeric histone H3 antibody in Allium
Abstract
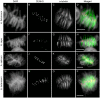
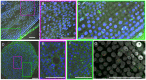
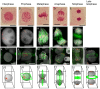
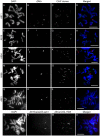
Due to the ease with which chromosomes can be observed, the Allium species, and onion in particular, have been familiar materials employed in cytogenetic experiments in biology. In this study, centromeric histone H3 (CENH3)-coding cDNAs were identified in four Allium species (onion, welsh onion, garlic and garlic chives) and cloned. Anti-CENH3 antibody was then raised against a deduced amino acid sequence of CENH3 of welsh onion. The antibody recognized all CENH3 orthologs of the Allium species tested. Immunostaining with the antibody enabled clear visualization of chromosome behavior during mitosis in the species. Furthermore, three-dimensional (3D) observation of mitotic cell division was achieved by subjecting root sections to immunohistochemical techniques. The 3D dynamics of the cells and position of cell-cycle marker proteins (CENH3 and α-tubulin) were clearly revealed by immunohistochemical staining with the antibodies. The immunohistochemical analysis made it possible to establish an overview of the location of dividing cells in the root tissues. This breakthrough in technique, in addition to the two centromeric DNA sequences isolated from welsh onion by chromatin immuno-precipitation using the antibody, should lead to a better understanding of plant cell division. A phylogenetic analysis of Allium CENH3s together with the previously reported plant CENH3s showed two separate clades for monocot species tested. One clade was made from CENH3s of the Allium species with those of Poaceae species, and the other from CENH3s of a holocentric species (Luzula nivea). These data may imply functional differences of CENH3s between holocentric and monocentric species. Centromeric localization of DNA sequences isolated from welsh onion by chromatin immuno-precipitation (ChIP) using the antibody was confirmed by fluorescence in situ hybridization and ChIP-quantitative PCR.
Conflict of interest statement
Figures

Similar articles
-
Pancentromere analysis of Allium species reveals diverse centromere positions in onion and gigantic centromeres in garlic.Plant Cell. 2025 Jul 1;37(7):koaf142. doi: 10.1093/plcell/koaf142. Plant Cell. 2025. PMID: 40493742 Free PMC article.
-
Visualization of diffuse centromeres with centromere-specific histone H3 in the holocentric plant Luzula nivea.Plant Cell. 2005 Jul;17(7):1886-93. doi: 10.1105/tpc.105.032961. Epub 2005 Jun 3. Plant Cell. 2005. PMID: 15937225 Free PMC article.
-
Characterization of centromeric histone H3 (CENH3) variants in cultivated and wild carrots (Daucus sp.).PLoS One. 2014 Jun 2;9(6):e98504. doi: 10.1371/journal.pone.0098504. eCollection 2014. PLoS One. 2014. PMID: 24887084 Free PMC article.
-
Centromere dynamics.Curr Opin Genet Dev. 2007 Apr;17(2):151-6. doi: 10.1016/j.gde.2007.02.009. Epub 2007 Feb 22. Curr Opin Genet Dev. 2007. PMID: 17320374 Review.
-
Allium cytogenetics: a critical review on the Indian taxa.Comp Cytogenet. 2023 May 29;17:129-156. doi: 10.3897/CompCytogen.17.98903. eCollection 2023. Comp Cytogenet. 2023. PMID: 37304149 Free PMC article. Review.
Cited by
-
Chromosomal variations of Lycoris species revealed by FISH with rDNAs and centromeric histone H3 variant associated DNAs.PLoS One. 2021 Sep 30;16(9):e0258028. doi: 10.1371/journal.pone.0258028. eCollection 2021. PLoS One. 2021. PMID: 34591908 Free PMC article.
-
Tandem repeats of Allium fistulosum associated with major chromosomal landmarks.Mol Genet Genomics. 2017 Apr;292(2):453-464. doi: 10.1007/s00438-016-1286-9. Epub 2017 Feb 1. Mol Genet Genomics. 2017. PMID: 28150039
-
Pancentromere analysis of Allium species reveals diverse centromere positions in onion and gigantic centromeres in garlic.Plant Cell. 2025 Jul 1;37(7):koaf142. doi: 10.1093/plcell/koaf142. Plant Cell. 2025. PMID: 40493742 Free PMC article.
-
Immuno-cytogenetic manifestation of epigenetic chromatin modification marks in plants.Planta. 2015 Feb;241(2):291-301. doi: 10.1007/s00425-014-2233-9. Epub 2014 Dec 25. Planta. 2015. PMID: 25539867
-
Comparative Dissection of Three Giant Genomes: Allium cepa, Allium sativum, and Allium ursinum.Int J Mol Sci. 2019 Feb 9;20(3):733. doi: 10.3390/ijms20030733. Int J Mol Sci. 2019. PMID: 30744119 Free PMC article.
References
-
- Leme DM, Marin-Morales MA (2009) Allium cepa test in environmental monitoring: a review on its application. Mutat Res 682: 71–81. - PubMed
-
- Alberts B, Johnson A, Lewis J, Raff M, Roberts K, et al... (2007) Molecular Biology of the Cell; 5, editor. New York: Garland Science.
-
- Simon EJ, Reece JB, Dickey JL (2010) Campbell Essential Biology with Physiology: Benjamin Cummings.
-
- Lodish H, Berk A, Kaiser CA, Krieger M, Scott MP, et al... (2007) Molecular Cell Biology: W. H. Freeman.
-
- Pich U, Fuchs J, Schubert I (1996) How do Alliaceae stabilize their chromosome ends in the absence of TTTAGGG sequences? Chromosome Res 4: 207–213. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

