Genetic variability among complete human respiratory syncytial virus subgroup A genomes: bridging molecular evolutionary dynamics and epidemiology
- PMID: 23236501
- PMCID: PMC3517519
- DOI: 10.1371/journal.pone.0051439
Genetic variability among complete human respiratory syncytial virus subgroup A genomes: bridging molecular evolutionary dynamics and epidemiology
Abstract
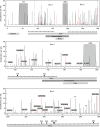
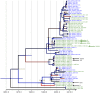
Human respiratory syncytial virus (RSV) is an important cause of severe lower respiratory tract infections in infants and the elderly. In the vast majority of cases, however, RSV infections run mild and symptoms resemble those of a common cold. The immunological, clinical, and epidemiological profile of severe RSV infections suggests a disease caused by a virus with typical seasonal transmission behavior, lacking clear-cut virulence factors, but instead causing disease by modifying the host's immune response in a way that stimulates pathogenesis. Yet, the interplay between RSV-evoked immune responses and epidemic behavior, and how this affects the genomic evolutionary dynamics of the virus, remains poorly understood. Here, we present a comprehensive collection of 33 novel RSV subgroup A genomes from strains sampled over the last decade, and provide the first measurement of RSV-A genomic diversity through time in a phylodynamic framework. In addition, we map amino acid substitutions per protein to determine mutational hotspots in specific domains. Using Bayesian genealogical inference, we estimated the genomic evolutionary rate to be 6.47 × 10(-4) (credible interval: 5.56 × 10(-4), 7.38 × 10(-4)) substitutions/site/year, considerably slower than previous estimates based on G gene sequences only. The G gene is however marked by elevated substitution rates compared to other RSV genes, which can be attributed to relaxed selective constraints. In line with this, site-specific selection analyses identify the G gene as the major target of diversifying selection. Importantly, statistical analysis demonstrates that the immune driven positive selection does not leave a measurable imprint on the genome phylogeny, implying that RSV lineage replacement mainly follows nonselective epidemiological processes. The roughly 50 years of RSV-A genomic evolution are characterized by a constant population size through time and general co-circulation of lineages over many epidemic seasons - a conclusion that might be taken into account when developing future therapeutic and preventive strategies.
Conflict of interest statement
Figures



References
-
- Glezen WP, Taber LH, Frank AL, Kasel JA (1986) Risk of primary infection and reinfection with respiratory syncytial virus. Am J Dis Child 140: 543–546. - PubMed
-
- Henrickson KJ, Hoover S, Kehl KS, Hua W (2004) National disease burden of respiratory viruses detected in children by polymerase chain reaction. Pediatr Infect Dis J 23: S11–18. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

