Engineering of Corynebacterium glutamicum for high-yield L-valine production under oxygen deprivation conditions
- PMID: 23241971
- PMCID: PMC3568611
- DOI: 10.1128/AEM.02806-12
Engineering of Corynebacterium glutamicum for high-yield L-valine production under oxygen deprivation conditions
Abstract
We previously demonstrated efficient L-valine production by metabolically engineered Corynebacterium glutamicum under oxygen deprivation. To achieve the high productivity, a NADH/NADPH cofactor imbalance during the synthesis of l-valine was overcome by engineering NAD-preferring mutant acetohydroxy acid isomeroreductase (AHAIR) and using NAD-specific leucine dehydrogenase from Lysinibacillus sphaericus. Lactate as a by-product was largely eliminated by disrupting the lactate dehydrogenase gene ldhA. Nonetheless, a few other by-products, particularly succinate, were still produced and acted to suppress the L-valine yield. Eliminating these by-products therefore was deemed key to improving theL-valine yield. By additionally disrupting the phosphoenolpyruvate carboxylase gene ppc, succinate production was effectively suppressed, but both glucose consumption and L-valine production dropped considerably due to the severely elevated intracellular NADH/NAD(+) ratio. In contrast, this perturbed intracellular redox state was more than compensated for by deletion of three genes associated with NADH-producing acetate synthesis and overexpression of five glycolytic genes, including gapA, encoding NADH-inhibited glyceraldehyde-3-phosphate dehydrogenase. Inserting feedback-resistant mutant acetohydroxy acid synthase and NAD-preferring mutant AHAIR in the chromosome resulted in higher L-valine yield and productivity. Deleting the alanine transaminase gene avtA suppressed alanine production. The resultant strain produced 1,280 mM L-valine at a yield of 88% mol mol of glucose(-1) after 24 h under oxygen deprivation, a vastly improved yield over our previous best.
Figures

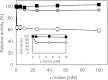
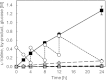
References
-
- Park JH, Lee SY. 2010. Fermentative production of branched chain amino acids: a focus on metabolic engineering. Appl. Microbiol. Biotechnol. 85:491–506 - PubMed
-
- Hermann T. 2003. Industrial production of amino acids by coryneform bacteria. J. Biotechnol. 104:155–172 - PubMed
-
- Blombach B, Schreiner ME, Bartek T, Oldiges M, Eikmanns BJ. 2008. Corynebacterium glutamicum tailored for high-yield L-valine production. Appl. Microbiol. Biotechnol. 79:471–479 - PubMed
-
- Durre P. 2007. Biobutanol: an attractive biofuel. Biotechnol. J. 2:1525–1534 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

