Matrix assisted ionization vacuum (MAIV), a new ionization method for biological materials analysis using mass spectrometry
- PMID: 23242551
- PMCID: PMC3591669
- DOI: 10.1074/mcp.M112.023663
Matrix assisted ionization vacuum (MAIV), a new ionization method for biological materials analysis using mass spectrometry
Abstract
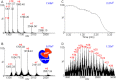
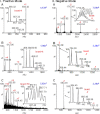
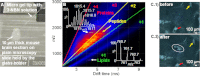
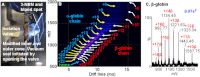
The introduction of electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI) for the mass spectrometric analysis of peptides and proteins had a dramatic impact on biological science. We now report that a wide variety of compounds, including peptides, proteins, and protein complexes, are transported directly from a solid-state small molecule matrix to gas-phase ions when placed into the vacuum of a mass spectrometer without the use of high voltage, a laser, or added heat. This ionization process produces ions having charge states similar to ESI, making the method applicable for high performance mass spectrometers designed for atmospheric pressure ionization. We demonstrate highly sensitive ionization using intermediate pressure MALDI and modified ESI sources. This matrix and vacuum assisted soft ionization method is suitable for the direct surface analysis of biological materials, including tissue, via mass spectrometry.
Figures
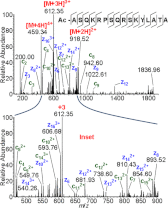
References
-
- Fenn J. B., Mann M., Meng C. K., Wong S. F., Whitehouse C. M. (1989) Electrospray ionization for mass spectrometry of large biomolecules. Science 246, 64–71 - PubMed
-
- Tanaka K., Waki H., Ido Y., Akita S., Yoshida Y. (1988) Protein and polymer analyses up to m/z 100 000 by laser ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 2, 151–153
-
- Vestal M. L. (2009) Modern MALDI time-of-flight mass spectrometry. J. Mass Spectrom. 44, 303–317 - PubMed
-
- Rosinke B., Strupat K., Hillenkamp F., Rosenbusch J., Dencher N., Kruger U., Galla H. J. (1995) Matrix-assisted laser desorption/ionization mass-spectrometry (MALDI-MS) of membrane-proteins and noncovalent complexes. J. Mass Spectrom. 30, 1462–1468
-
- Bich C., Zenobi R. (2009) Mass spectrometry of large complexes. Curr. Opin. Struct. Biol. 19, 632–639 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

