Sculpting of DNA at abasic sites by DNA glycosylase homolog mag2
- PMID: 23245849
- PMCID: PMC3545110
- DOI: 10.1016/j.str.2012.11.004
Sculpting of DNA at abasic sites by DNA glycosylase homolog mag2
Abstract
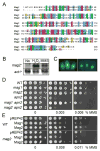
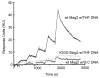
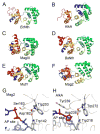
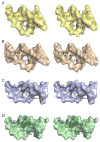
Modifications and loss of bases are frequent types of DNA lesions, often handled by the base excision repair (BER) pathway. BER is initiated by DNA glycosylases, generating abasic (AP) sites that are subsequently cleaved by AP endonucleases, which further pass on nicked DNA to downstream DNA polymerases and ligases. The coordinated handover of cytotoxic intermediates between different BER enzymes is most likely facilitated by the DNA conformation. Here, we present the atomic structure of Schizosaccharomyces pombe Mag2 in complex with DNA to reveal an unexpected structural basis for nonenzymatic AP site recognition with an unflipped AP site. Two surface-exposed loops intercalate and widen the DNA minor groove to generate a DNA conformation previously only found in the mismatch repair MutS-DNA complex. Consequently, the molecular role of Mag2 appears to be AP site recognition and protection, while possibly facilitating damage signaling by structurally sculpting the DNA substrate.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures



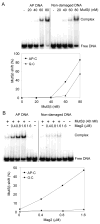
References
-
- Berti PJ, McCann JA. Toward a detailed understanding of base excision repair enzymes: transition state and mechanistic analyses of N-glycoside hydrolysis and N-glycoside transfer. Chem Rev. 2006;106:506–555. - PubMed
-
- Boiteux S, Guillet M. Abasic sites in DNA: repair and biological consequences in Saccharomyces cerevisiae. DNA Repair (Amst) 2004;3:1–12. - PubMed
-
- Bruner SD, Norman DP, Verdine GL. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature. 2000;403:859–866. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

