Parallel labeling experiments and metabolic flux analysis: Past, present and future methodologies
- PMID: 23246523
- PMCID: PMC6474253
- DOI: 10.1016/j.ymben.2012.11.010
Parallel labeling experiments and metabolic flux analysis: Past, present and future methodologies
Abstract
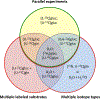
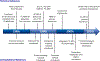
Radioactive and stable isotopes have been applied for decades to elucidate metabolic pathways and quantify carbon flow in cellular systems using mass and isotope balancing approaches. Isotope-labeling experiments can be conducted as a single tracer experiment, or as parallel labeling experiments. In the latter case, several experiments are performed under identical conditions except for the choice of substrate labeling. In this review, we highlight robust approaches for probing metabolism and addressing metabolically related questions though parallel labeling experiments. In the first part, we provide a brief historical perspective on parallel labeling experiments, from the early metabolic studies when radioisotopes were predominant to present-day applications based on stable-isotopes. We also elaborate on important technical and theoretical advances that have facilitated the transition from radioisotopes to stable-isotopes. In the second part of the review, we focus on parallel labeling experiments for (13)C-metabolic flux analysis ((13)C-MFA). Parallel experiments offer several advantages that include: tailoring experiments to resolve specific fluxes with high precision; reducing the length of labeling experiments by introducing multiple entry-points of isotopes; validating biochemical network models; and improving the performance of (13)C-MFA in systems where the number of measurements is limited. We conclude by discussing some challenges facing the use of parallel labeling experiments for (13)C-MFA and highlight the need to address issues related to biological variability, data integration, and rational tracer selection.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Integrated 13C-metabolic flux analysis of 14 parallel labeling experiments in Escherichia coli.Metab Eng. 2015 Mar;28:151-158. doi: 10.1016/j.ymben.2015.01.001. Epub 2015 Jan 14. Metab Eng. 2015. PMID: 25596508 Free PMC article.
-
COMPLETE-MFA: complementary parallel labeling experiments technique for metabolic flux analysis.Metab Eng. 2013 Nov;20:49-55. doi: 10.1016/j.ymben.2013.08.006. Epub 2013 Sep 8. Metab Eng. 2013. PMID: 24021936
-
Parallel labeling experiments with [U-13C]glucose validate E. coli metabolic network model for 13C metabolic flux analysis.Metab Eng. 2012 Sep;14(5):533-41. doi: 10.1016/j.ymben.2012.06.003. Epub 2012 Jul 6. Metab Eng. 2012. PMID: 22771935
-
13C metabolic flux analysis: optimal design of isotopic labeling experiments.Curr Opin Biotechnol. 2013 Dec;24(6):1116-21. doi: 10.1016/j.copbio.2013.02.003. Epub 2013 Feb 28. Curr Opin Biotechnol. 2013. PMID: 23453397 Review.
-
Isotopically nonstationary metabolic flux analysis (INST-MFA): putting theory into practice.Curr Opin Biotechnol. 2018 Dec;54:80-87. doi: 10.1016/j.copbio.2018.02.013. Epub 2018 Mar 6. Curr Opin Biotechnol. 2018. PMID: 29522915 Review.
Cited by
-
Isotopically nonstationary 13C metabolic flux analysis in resting and activated human platelets.Metab Eng. 2022 Jan;69:313-322. doi: 10.1016/j.ymben.2021.12.007. Epub 2021 Dec 22. Metab Eng. 2022. PMID: 34954086 Free PMC article.
-
Co-utilization of glucose and xylose by evolved Thermus thermophilus LC113 strain elucidated by (13)C metabolic flux analysis and whole genome sequencing.Metab Eng. 2016 Sep;37:63-71. doi: 10.1016/j.ymben.2016.05.001. Epub 2016 May 7. Metab Eng. 2016. PMID: 27164561 Free PMC article.
-
Metabolic flux analysis of Escherichia coli knockouts: lessons from the Keio collection and future outlook.Curr Opin Biotechnol. 2014 Aug;28:127-33. doi: 10.1016/j.copbio.2014.02.006. Epub 2014 Mar 28. Curr Opin Biotechnol. 2014. PMID: 24686285 Free PMC article. Review.
-
Comprehensive metabolic modeling of multiple 13C-isotopomer data sets to study metabolism in perfused working hearts.Am J Physiol Heart Circ Physiol. 2016 Oct 1;311(4):H881-H891. doi: 10.1152/ajpheart.00428.2016. Epub 2016 Aug 5. Am J Physiol Heart Circ Physiol. 2016. PMID: 27496880 Free PMC article.
-
OpenFLUX2: (13)C-MFA modeling software package adjusted for the comprehensive analysis of single and parallel labeling experiments.Microb Cell Fact. 2014 Nov 19;13:152. doi: 10.1186/s12934-014-0152-x. Microb Cell Fact. 2014. PMID: 25408234 Free PMC article.
References
-
- Ahn WS, Antoniewicz MR, 2011. Metabolic flux analysis of CHO cells at growth and non-growth phases using isotopic tracers and mass spectrometry. Metab Eng. 13, 598–609. - PubMed
-
- Ahn WS, Antoniewicz MR, 2012. Towards dynamic metabolic flux analysis in CHO cell cultures. Biotechnol J. 7, 61–74. - PubMed
-
- Allen DK, Ohlrogge JB, Shachar-Hill Y, 2009. The role of light in soybean seed filling metabolism. Plant J. 58, 220–34. - PubMed
-
- Alonso AP, Dale VL, Shachar-Hill Y, 2010. Understanding fatty acid synthesis in developing maize embryos using metabolic flux analysis. Metab Eng. 12, 488–97. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

