Competing and noncompeting activities of miR-122 and the 5' exonuclease Xrn1 in regulation of hepatitis C virus replication
- PMID: 23248316
- PMCID: PMC3562843
- DOI: 10.1073/pnas.1213515110
Competing and noncompeting activities of miR-122 and the 5' exonuclease Xrn1 in regulation of hepatitis C virus replication
Abstract
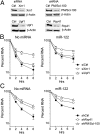
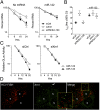
Hepatitis C virus (HCV) replication is dependent on microRNA 122 (miR-122), a liver-specific microRNA that recruits Argonaute 2 to the 5' end of the viral genome, stabilizing it and slowing its decay both in cell-free reactions and in infected cells. Here we describe the RNA degradation pathways against which miR-122 provides protection. Transfected HCV RNA is degraded by both the 5' exonuclease Xrn1 and 3' exonuclease exosome complex, whereas replicating RNA within infected cells is degraded primarily by Xrn1 with no contribution from the exosome. Consistent with this, sequencing of the 5' and 3' ends of RNA degradation intermediates in infected cells confirmed that 5' decay is the primary pathway for HCV RNA degradation. Xrn1 knockdown enhances HCV replication, indicating that Xrn1 decay and the viral replicase compete to set RNA abundance within infected cells. Xrn1 knockdown and miR-122 supplementation have equal, redundant, and nonadditive effects on the rate of viral RNA decay, indicating that miR-122 protects HCV RNA from 5' decay. Nevertheless, Xrn1 knockdown does not rescue replication of a viral mutant defective in miR-122 binding, indicating that miR-122 has additional yet uncharacterized function(s) in the viral life cycle.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Comment in
-
miR-122 is more than a shield for the hepatitis C virus genome.Proc Natl Acad Sci U S A. 2013 Jan 29;110(5):1571-2. doi: 10.1073/pnas.1220841110. Epub 2013 Jan 16. Proc Natl Acad Sci U S A. 2013. PMID: 23324744 Free PMC article. No abstract available.
References
-
- Lemon SM, Walker C, Alter MJ, Yi M. Hepatitis C viruses. In: Knipe DM, et al., editors. Fields Virology. 5th Ed. Philadelphia: Lippincott Williams & Wilkins; 2007. pp. 1253–1304.
-
- Chang J, et al. miR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol. 2004;1(2):106–113. - PubMed
-
- Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science. 2005;309(5740):1577–1581. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

