A time-calibrated molecular phylogeny of the precious corals: reconciling discrepancies in the taxonomic classification and insights into their evolutionary history
- PMID: 23249327
- PMCID: PMC3565870
- DOI: 10.1186/1471-2148-12-246
A time-calibrated molecular phylogeny of the precious corals: reconciling discrepancies in the taxonomic classification and insights into their evolutionary history
Abstract
Background: Seamount-associated faunas are often considered highly endemic but isolation and diversification processes leading to such endemism have been poorly documented at those depths. Likewise, species delimitation and phylogenetic studies in deep-sea organisms remain scarce, due to the difficulty in obtaining samples, and sometimes controversial. The phylogenetic relationships within the precious coral family Coralliidae remain largely unexplored and the monophyly of its two constituent genera, Corallium Cuvier and Paracorallium Bayer & Cairns, has not been resolved. As traditionally recognized, the diversity of colonial forms among the various species correlates with the diversity in shape of their supporting axis, but the phylogenetic significance of these characters remains to be tested. We thus used mitochondrial sequence data to evaluate the monophyly of Corallium and Paracorallium and the species boundaries for nearly all named taxa in the family. Species from across the coralliid range, including material from Antarctica, Hawaii, Japan, New Zealand, Taiwan, Tasmania, the eastern Pacific and the western Atlantic were examined.
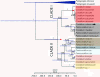
Results: The concatenated analysis of five mitochondrial regions (COI, 16S rRNA, ND2, and ND3-ND6) recovered two major coralliid clades. One clade is composed of two subgroups, the first including Corallium rubrum, the type species of the genus, together with a small group of Paracorallium species (P. japonicum and P. tortuosum) and C. medea (clade I-A); the other subgroup includes a poorly-resolved assemblage of six Corallium species (C. abyssale, C. ducale, C. imperiale, C. laauense, C. niobe, and C. sulcatum; clade I-B). The second major clade is well resolved and includes species of Corallium and Paracorallium (C. elatius, C. kishinouyei, C. konojoi, C. niveum, C. secundum, Corallium sp., Paracorallium nix, Paracorallium thrinax and Paracorallium spp.). A traditional taxonomic study of this clade delineated 11 morphospecies that were congruent with the general mixed Yule-coalescent (GMYC) model. A multilocus species-tree approach also identified the same two well-supported clades, being Clade I-B more recent in the species tree (18.0-15.9 mya) than in the gene tree (35.2-15.9 mya). In contrast, the diversification times for Clade II were more ancient in the species tree (136.4-41.7 mya) than in the gene tree (66.3-16.9 mya).
Conclusions: Our results provide no support for the taxonomic status of the two currently recognized genera in the family Coralliidae. Given that Paracorallium species were all nested within Corallium, we recognize the coralliid genus Corallium, which includes the type species of the family, and thus consider Paracorallium a junior synonym of Corallium. We propose the use of the genus Hemicorallium Gray for clade I-B (species with long rod sclerites, cylindrical autozooids and smooth axis). Species delimitation in clade I-B remains unclear and the molecular resolution for Coralliidae species is inconsistent in the two main clades. Some species have wide distributions, recent diversification times and low mtDNA divergence whereas other species exhibit narrower allopatric distributions, older diversification times and greater levels of mtDNA resolution.
Figures



References
-
- Lim GS, Balke M, Meier R. Determining species boundaries in a world full of rarity: singletons, species delimitation methods. Syst Biol. 2011;60:165–169. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

