Differences in enhancer activity in mouse and zebrafish reporter assays are often associated with changes in gene expression
- PMID: 23253453
- PMCID: PMC3541358
- DOI: 10.1186/1471-2164-13-713
Differences in enhancer activity in mouse and zebrafish reporter assays are often associated with changes in gene expression
Abstract
Background: Phenotypic evolution in animals is thought to be driven in large part by differences in gene expression patterns, which can result from sequence changes in cis-regulatory elements (cis-changes) or from changes in the expression pattern or function of transcription factors (trans-changes). While isolated examples of trans-changes have been identified, the scale of their overall contribution to regulatory and phenotypic evolution remains unclear.
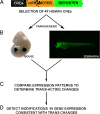
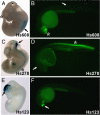
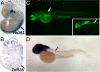
Results: Here, we attempt to examine the prevalence of trans-effects and their potential impact on gene expression patterns in vertebrate evolution by comparing the function of identical human tissue-specific enhancer sequences in two highly divergent vertebrate model systems, mouse and zebrafish. Among 47 human conserved non-coding elements (CNEs) tested in transgenic mouse embryos and in stable zebrafish lines, at least one species-specific expression domain was observed in the majority (83%) of cases, and 36% presented dramatically different expression patterns between the two species. Although some of these discrepancies may be due to the use of different transgenesis systems in mouse and zebrafish, in some instances we found an association between differences in enhancer activity and changes in the endogenous gene expression patterns between mouse and zebrafish, suggesting a potential role for trans-changes in the evolution of gene expression.
Conclusions: In total, our results: (i) serve as a cautionary tale for studies investigating the role of human enhancers in different model organisms, and (ii) suggest that changes in the trans environment may play a significant role in the evolution of gene expression in vertebrates.
Figures
Similar articles
-
The importance of being cis: evolution of orthologous fish and mammalian enhancer activity.Mol Biol Evol. 2010 Oct;27(10):2322-32. doi: 10.1093/molbev/msq128. Epub 2010 May 21. Mol Biol Evol. 2010. PMID: 20494938 Free PMC article.
-
Transgenic analysis of Dlx regulation in fish tooth development reveals evolutionary retention of enhancer function despite organ loss.Proc Natl Acad Sci U S A. 2006 Dec 19;103(51):19390-5. doi: 10.1073/pnas.0609575103. Epub 2006 Dec 4. Proc Natl Acad Sci U S A. 2006. PMID: 17146045 Free PMC article.
-
Cis-regulatory characterization of sequence conservation surrounding the Hox4 genes.Dev Biol. 2010 Apr 15;340(2):269-82. doi: 10.1016/j.ydbio.2010.01.035. Epub 2010 Feb 6. Dev Biol. 2010. PMID: 20144609
-
Retroviral enhancer detection insertions in zebrafish combined with comparative genomics reveal genomic regulatory blocks - a fundamental feature of vertebrate genomes.Genome Biol. 2007;8 Suppl 1(Suppl 1):S4. doi: 10.1186/gb-2007-8-s1-s4. Genome Biol. 2007. PMID: 18047696 Free PMC article. Review.
-
Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence.Nat Rev Genet. 2011 Dec 6;13(1):59-69. doi: 10.1038/nrg3095. Nat Rev Genet. 2011. PMID: 22143240 Review.
Cited by
-
Genomic perspectives of transcriptional regulation in forebrain development.Neuron. 2015 Jan 7;85(1):27-47. doi: 10.1016/j.neuron.2014.11.011. Neuron. 2015. PMID: 25569346 Free PMC article. Review.
-
Exaptation of transposable elements into novel cis-regulatory elements: is the evidence always strong?Mol Biol Evol. 2013 Jun;30(6):1239-51. doi: 10.1093/molbev/mst045. Epub 2013 Mar 13. Mol Biol Evol. 2013. PMID: 23486611 Free PMC article. Review.
-
Twenty-Seven Tamoxifen-Inducible iCre-Driver Mouse Strains for Eye and Brain, Including Seventeen Carrying a New Inducible-First Constitutive-Ready Allele.Genetics. 2019 Apr;211(4):1155-1177. doi: 10.1534/genetics.119.301984. Epub 2019 Feb 14. Genetics. 2019. PMID: 30765420 Free PMC article.
-
Biocompatibility Analysis of Bio-Based and Synthetic Silica Nanoparticles during Early Zebrafish Development.Int J Mol Sci. 2024 May 18;25(10):5530. doi: 10.3390/ijms25105530. Int J Mol Sci. 2024. PMID: 38791566 Free PMC article.
-
Enhancer turnover and conserved regulatory function in vertebrate evolution.Philos Trans R Soc Lond B Biol Sci. 2013 Nov 11;368(1632):20130027. doi: 10.1098/rstb.2013.0027. Print 2013 Dec 19. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 24218639 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

