Pyruvate:ferredoxin oxidoreductase is coupled to light-independent hydrogen production in Chlamydomonas reinhardtii
- PMID: 23258532
- PMCID: PMC3567687
- DOI: 10.1074/jbc.M112.429985
Pyruvate:ferredoxin oxidoreductase is coupled to light-independent hydrogen production in Chlamydomonas reinhardtii
Abstract
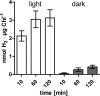
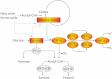
In anaerobiosis, the green alga Chlamydomonas reinhardtii evolves molecular hydrogen (H(2)) as one of several fermentation products. H(2) is generated mostly by the [Fe-Fe]-hydrogenase HYDA1, which uses plant type ferredoxin PETF/FDX1 (PETF) as an electron donor. Dark fermentation of the alga is mainly of the mixed acid type, because formate, ethanol, and acetate are generated by a pyruvate:formate lyase pathway similar to Escherichia coli. However, C. reinhardtii also possesses the pyruvate:ferredoxin oxidoreductase PFR1, which, like pyruvate:formate lyase and HYDA1, is localized in the chloroplast. PFR1 has long been suggested to be responsible for the low but significant H(2) accumulation in the dark because the catalytic mechanism of pyruvate:ferredoxin oxidoreductase involves the reduction of ferredoxin. With the aim of proving the biochemical feasibility of the postulated reaction, we have heterologously expressed the PFR1 gene in E. coli. Purified recombinant PFR1 is able to transfer electrons from pyruvate to HYDA1, using the ferredoxins PETF and FDX2 as electron carriers. The high reactivity of PFR1 toward oxaloacetate indicates that in vivo, fermentation might also be coupled to an anaerobically active glyoxylate cycle. Our results suggest that C. reinhardtii employs a clostridial type H(2) production pathway in the dark, especially because C. reinhardtii PFR1 was also able to allow H(2) evolution in reaction mixtures containing Clostridium acetobutylicum 2[4Fe-4S]-ferredoxin and [Fe-Fe]-hydrogenase HYDA.
Figures




Similar articles
-
Chlamydomonas reinhardtii chloroplasts contain a homodimeric pyruvate:ferredoxin oxidoreductase that functions with FDX1.Plant Physiol. 2013 Jan;161(1):57-71. doi: 10.1104/pp.112.208181. Epub 2012 Nov 15. Plant Physiol. 2013. PMID: 23154536 Free PMC article.
-
Crystal structure and biochemical characterization of Chlamydomonas FDX2 reveal two residues that, when mutated, partially confer FDX2 the redox potential and catalytic properties of FDX1.Photosynth Res. 2016 Apr;128(1):45-57. doi: 10.1007/s11120-015-0198-6. Epub 2015 Nov 3. Photosynth Res. 2016. PMID: 26526668 Free PMC article.
-
A novel, anaerobically induced ferredoxin in Chlamydomonas reinhardtii.FEBS Lett. 2009 Jan 22;583(2):325-9. doi: 10.1016/j.febslet.2008.12.018. Epub 2008 Dec 26. FEBS Lett. 2009. PMID: 19101555
-
Multiple ferredoxin isoforms in Chlamydomonas reinhardtii - their role under stress conditions and biotechnological implications.Eur J Cell Biol. 2010 Dec;89(12):998-1004. doi: 10.1016/j.ejcb.2010.06.018. Epub 2010 Aug 8. Eur J Cell Biol. 2010. PMID: 20696493 Review.
-
Evolution of Chlamydomonas reinhardtii ferredoxins and their interactions with [FeFe]-hydrogenases.Photosynth Res. 2017 Dec;134(3):307-316. doi: 10.1007/s11120-017-0409-4. Epub 2017 Jun 15. Photosynth Res. 2017. PMID: 28620699 Review.
Cited by
-
Critical role of Chlamydomonas reinhardtii ferredoxin-5 in maintaining membrane structure and dark metabolism.Proc Natl Acad Sci U S A. 2015 Dec 1;112(48):14978-83. doi: 10.1073/pnas.1515240112. Epub 2015 Nov 16. Proc Natl Acad Sci U S A. 2015. PMID: 26627249 Free PMC article.
-
Changing the tracks: screening for electron transfer proteins to support hydrogen production.J Biol Inorg Chem. 2022 Oct;27(7):631-640. doi: 10.1007/s00775-022-01956-1. Epub 2022 Aug 29. J Biol Inorg Chem. 2022. PMID: 36038787 Free PMC article.
-
Designing a modified clostridial 2[4Fe-4S] ferredoxin as a redox coupler to directly link photosystem I with a Pt nanoparticle.Photosynth Res. 2020 Feb;143(2):165-181. doi: 10.1007/s11120-019-00679-w. Epub 2019 Oct 23. Photosynth Res. 2020. PMID: 31643016
-
Engineering photosynthetic organisms for the production of biohydrogen.Photosynth Res. 2015 Mar;123(3):241-53. doi: 10.1007/s11120-014-9991-x. Epub 2014 Mar 27. Photosynth Res. 2015. PMID: 24671643 Free PMC article. Review.
-
On the pathways feeding the H2 production process in nutrient-replete, hypoxic conditions. Commentary on the article "Low oxygen levels contribute to improve photohydrogen production in mixotrophic non-stressed Chlamydomonas cultures", by Jurado-Oller et al., Biotechnology for Biofuels, published September 7, 2015; 8:149.Biotechnol Biofuels. 2017 May 4;10:116. doi: 10.1186/s13068-017-0800-6. eCollection 2017. Biotechnol Biofuels. 2017. PMID: 28484516 Free PMC article.
References
-
- Hemschemeier A., Happe T. (2005) The exceptional photofermentative hydrogen metabolism of the green alga Chlamydomonas reinhardtii. Biochem. Soc. Trans. 33, 39–41 - PubMed
-
- Atteia A., van Lis R., Gelius-Dietrich G., Adrait A., Garin J., Joyard J., Rolland N., Martin W. (2006) Pyruvate formate-lyase and a novel route of eukaryotic ATP synthesis in Chlamydomonas mitochondria. J. Biol. Chem. 281, 9909–9918 - PubMed
-
- Atteia A., van Lis R., Tielens A. G., Martin W. F. (2013) Anaerobic energy metabolism in unicellular photosynthetic eukaryotes. Biochim. Biophys. Acta 1827, 210–223 - PubMed
-
- Hemschemeier A., Happe T. (2011) Alternative photosynthetic electron transport pathways during anaerobiosis in the green alga Chlamydomonas reinhardtii. Biochim. Biophys. Acta 1807, 919–926 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

